Asking the Assay Expert: A Conversation on Innovation with qPCR Pioneer Professor Stephen Bustin

Want to listen to this article for FREE?
Complete the form below to unlock access to ALL audio articles.
Read time: 5 minutes
Over the past 15 years qPCR (real time quantitative Polymerase Chain Reaction) has developed considerably and has helped drive many advances in in vitro diagnostics. Developing a reliable and sensitive qPCR assay, however, remains a challenging process. In this Q&A Stephen Bustin Professor of Molecular Biology at Anglia Ruskin University, Chelmsford, UK, and lead-author of the MIQE: Minimum Information for Publication of Quantitative Real-Time PCR Experiments, offers insights into the challenges facing qPCR practitioners and how these are overcome through extensive quality assurance and fast Eco 48 (PCRmax) instrumentation.
Introducing qPCR: The diagnostic technique of choice
qPCR combines rapid and accurate target amplification with ease of use and has become a crucial tool in modern genomics research. The concept of qPCR involves measuring amplification products using a fluorescent label. During amplification, a fluorescent dye binds either directly, e.g. SYBR Green, or indirectly, e.g. via a labeled hybridising probe, to accumulating DNA molecules. The fluorescence values are then recorded during each cycle of the amplification process. qPCR allows accurate quantification and detection of target sequences. Today qPCR is the technique of choice across translational life science areas, from genomics and agrigenomics through to forensic science and disease diagnostics.
Over the past 15 years advances in instrumentation have substantially reduced run times for qPCR and greatly simplified process development. However, developing reliable qPCR assays still involves a number of expert tasks, including instrument and probe set-up, primer selection and, most importantly, process validation. Moreover, practical issues such as the speed of a run and the importance of precise block uniformity during heating are critical to the efficiency and reliability of results and conclusions derived from qPCR experiments.
In 2009 the MIQE guidelines were released to help harmonise qPCR practices and ensure the reliability of published data [1]. Among the points listed are those relating to the precision, limit of detection and linear dynamic range of qPCR instrumentation. These features aim to ensure that the instrument is capable of generating data to the level of accuracy and reliability required for meaningful results to be obtained. Modern instruments, such as the Eco 48 Thermal Cycler (PCRmax), are now developed to make meeting these guidelines simpler while simultaneously decreasing the time and cost of qPCR.
PCRmax: What are the major challenges facing contemporary qPCR in diagnostics?
Prof Bustin: Surprisingly, perhaps the most challenging aspect of qPCR remains developing an assay that generates good, meaningful results. As qPCR systems have advanced, users have become more distanced from the raw data, which makes it easier to overlook unreliability of results. This means researchers are often unaware of the true efficiency of their PCR reaction. Without robust validation it is impossible to know what the assay is detecting, which affects the reliability of experimental data. In fact a few recent reports have suggested that up to 90% of papers published in the biomedical research arena are unreproducible for such reasons. This is a worrying reminder of the importance of fully understanding the qPCR process and is a force that drove the development of the MIQE guidelines.
PCRmax: How can users mitigate these risks and ensure their results are reliable?
Prof Bustin: Robust quality assessment before progressing to sample analysis remains the most important aspect of method development. We spend a lot of time designing assays and making sure that everything is correct in silico before using fast qPCR instruments to corroborate and optimise these data. Only when we have adequately completed this step do we move on to empirical validation of the assay. Naturally, robust quality assessment and validation is an intensive process so employing a qPCR instrument that delivers rapid amplification within the MIQE framework is vital.
PCRmax: How big an impact do you think having the correct instrument has?
Prof Bustin: Obviously it is very important to choose the right technology. There are a lot of instruments on the market for qPCR and some will be more effective than others for certain applications. Over the past few years our team has worked on setting up a molecular diagnostic laboratory based on extending techniques such as qPCR to novel fungal and bacterial diagnostics. So for us, and anyone developing assays, an essential attribute you really need in an instrument is speed.
For example we are currently working on a project to assess inter-operatively lymph nodes in patients with breast cancer. The sentinel lymph node is removed from the patient in theatre and rushed to the lab for RNA extraction. A reverse transcription (RT) - qPCR reaction is then performed to screen for markers indicative of metastasis. We use the Eco 48 Thermal Cycler for this and the majority of our assay development work because it performs qPCR so quickly without any drop off in accuracy and reproducibility. In this particular case our optimised run has reduced the time taken to perform the entire assay from extraction to analysis down to just 29 minutes.
PCRmax: Where do you see qPCR going in the future?
Prof Bustin: I edit two journals and it is surprising how many cases I come across where conventional end point PCR is still being used for assays which it was never intended for. The first problem with end point PCR is that, unlike qPCR, it is necessary to run a gel so there is always the potential for contamination. The second is that endpoint PCR is not quantitative. End point PCR has some value, in applications such as genotyping and SNP analysis but for many other applications, robust qPCR techniques have been developed that are more appropriate. Moreover, qPCR instruments, like the Eco 48 can be used with sample volumes as low as 5 µL which, combined with the efficiency of systems such as the Eco 48, means the cost of running qPCR is much reduced. In the short term I expect to see qPCR completely replace end point techniques, with the exception of certain specialist applications.
Further ahead, one of the biggest developments is the maturation of digital PCR technology. Digital PCR has become much easier to deploy and is a technique we are definitely keeping an eye on. However there are some clear limitations, not least of all the high cost of the instruments and the complexity of the process. For this reason instruments like the Eco 48 still have clear advantages; qPCR is not only fast and robust but the software makes handling the reaction very straightforward. Moreover as most qPCR assays involve looking at the relative expression of one message over another it simply is not necessary to have the absolute precision in the quantity of target copies generated by digital PCR. I cannot see digital PCR being taken out into the field to monitor pathogens, for instance, while there are already hand-held battery driven qPCR devices available to perform such experiments. Because of its ubiquity and huge adaptability in terms of instrumentation, reagents and application it is safe to say we will still be using qPCR in ten years’ time.
The technology behind the research
With sensitivity of detection down to a single copy and the world’s most thermally accurate block, the Eco 48 from PCRmax enables researchers to surpass all required MIQE guidelines and accelerate run time, with no compromise to efficiency.
This is achieved through a number of innovations, including novel block technology which delivers complete heating uniformity and rapid cycles in under 40 minutes, with the sensitivity to deliver ±0.1oC uniformity across the whole block instantly after every temperature change (Figure 1).
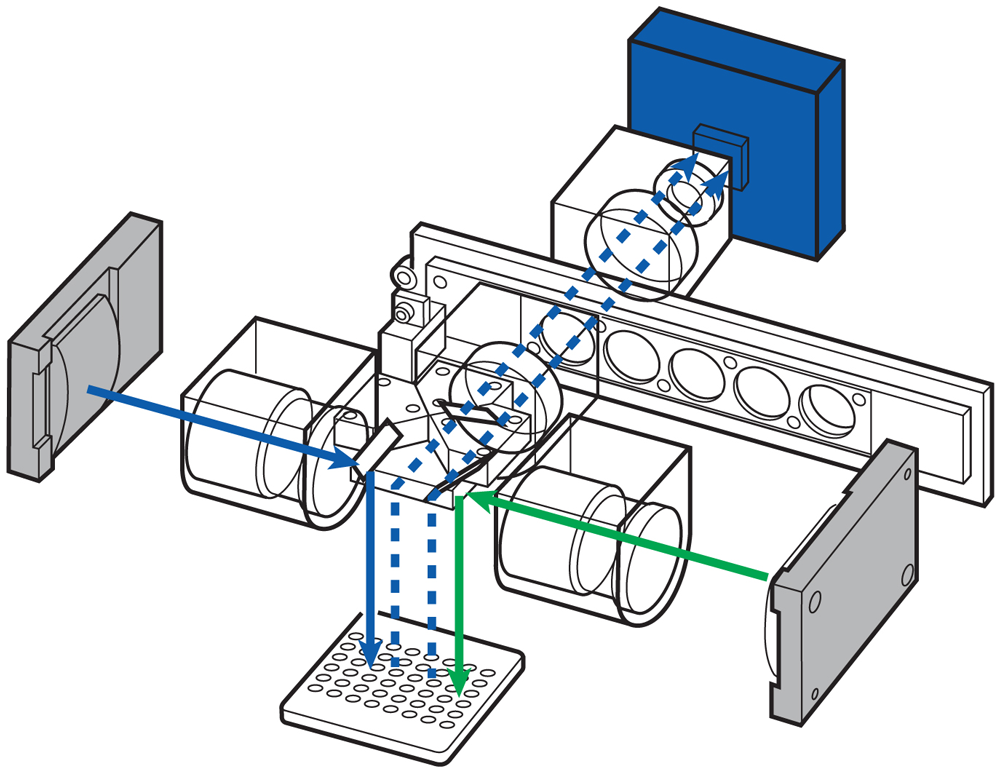
Meanwhile adaptive LED control a the high-performance optical system removes destructive variables from the fluorescence detection process, such as unknown saturation or light bleaching through to neighbouring cells (Figure 2).
Combined with intutive software, designed with ease of use in mind, the Eco 48 makes it faster and easier to develop rigorously validated assays that deliver complete confidence in results, improving the anaytical productivity for practically all qPCR application.
PCRmax is a Bibby Scientific company specialising in providing real time instrumentation, consumables and reagents for the life science laboratory. Find out more about PCRmax and the Eco 48 at www.pcrmax.com
For more information on Professor Bustin’s research contact enquiry@pcrmax.com
References
[1] Bustin S.A. et.al; The MIQE guidelines: minimum information for publication of quantitative real-time PCR experiments Clin Chem. 2009 Apr; 55(4):611-22
Introducing qPCR: The diagnostic technique of choice
qPCR combines rapid and accurate target amplification with ease of use and has become a crucial tool in modern genomics research. The concept of qPCR involves measuring amplification products using a fluorescent label. During amplification, a fluorescent dye binds either directly, e.g. SYBR Green, or indirectly, e.g. via a labeled hybridising probe, to accumulating DNA molecules. The fluorescence values are then recorded during each cycle of the amplification process. qPCR allows accurate quantification and detection of target sequences. Today qPCR is the technique of choice across translational life science areas, from genomics and agrigenomics through to forensic science and disease diagnostics.
Over the past 15 years advances in instrumentation have substantially reduced run times for qPCR and greatly simplified process development. However, developing reliable qPCR assays still involves a number of expert tasks, including instrument and probe set-up, primer selection and, most importantly, process validation. Moreover, practical issues such as the speed of a run and the importance of precise block uniformity during heating are critical to the efficiency and reliability of results and conclusions derived from qPCR experiments.

In 2009 the MIQE guidelines were released to help harmonise qPCR practices and ensure the reliability of published data [1]. Among the points listed are those relating to the precision, limit of detection and linear dynamic range of qPCR instrumentation. These features aim to ensure that the instrument is capable of generating data to the level of accuracy and reliability required for meaningful results to be obtained. Modern instruments, such as the Eco 48 Thermal Cycler (PCRmax), are now developed to make meeting these guidelines simpler while simultaneously decreasing the time and cost of qPCR.
PCRmax: What are the major challenges facing contemporary qPCR in diagnostics?
Prof Bustin: Surprisingly, perhaps the most challenging aspect of qPCR remains developing an assay that generates good, meaningful results. As qPCR systems have advanced, users have become more distanced from the raw data, which makes it easier to overlook unreliability of results. This means researchers are often unaware of the true efficiency of their PCR reaction. Without robust validation it is impossible to know what the assay is detecting, which affects the reliability of experimental data. In fact a few recent reports have suggested that up to 90% of papers published in the biomedical research arena are unreproducible for such reasons. This is a worrying reminder of the importance of fully understanding the qPCR process and is a force that drove the development of the MIQE guidelines.
PCRmax: How can users mitigate these risks and ensure their results are reliable?
Prof Bustin: Robust quality assessment before progressing to sample analysis remains the most important aspect of method development. We spend a lot of time designing assays and making sure that everything is correct in silico before using fast qPCR instruments to corroborate and optimise these data. Only when we have adequately completed this step do we move on to empirical validation of the assay. Naturally, robust quality assessment and validation is an intensive process so employing a qPCR instrument that delivers rapid amplification within the MIQE framework is vital.
PCRmax: How big an impact do you think having the correct instrument has?
Prof Bustin: Obviously it is very important to choose the right technology. There are a lot of instruments on the market for qPCR and some will be more effective than others for certain applications. Over the past few years our team has worked on setting up a molecular diagnostic laboratory based on extending techniques such as qPCR to novel fungal and bacterial diagnostics. So for us, and anyone developing assays, an essential attribute you really need in an instrument is speed.
For example we are currently working on a project to assess inter-operatively lymph nodes in patients with breast cancer. The sentinel lymph node is removed from the patient in theatre and rushed to the lab for RNA extraction. A reverse transcription (RT) - qPCR reaction is then performed to screen for markers indicative of metastasis. We use the Eco 48 Thermal Cycler for this and the majority of our assay development work because it performs qPCR so quickly without any drop off in accuracy and reproducibility. In this particular case our optimised run has reduced the time taken to perform the entire assay from extraction to analysis down to just 29 minutes.
PCRmax: Where do you see qPCR going in the future?
Prof Bustin: I edit two journals and it is surprising how many cases I come across where conventional end point PCR is still being used for assays which it was never intended for. The first problem with end point PCR is that, unlike qPCR, it is necessary to run a gel so there is always the potential for contamination. The second is that endpoint PCR is not quantitative. End point PCR has some value, in applications such as genotyping and SNP analysis but for many other applications, robust qPCR techniques have been developed that are more appropriate. Moreover, qPCR instruments, like the Eco 48 can be used with sample volumes as low as 5 µL which, combined with the efficiency of systems such as the Eco 48, means the cost of running qPCR is much reduced. In the short term I expect to see qPCR completely replace end point techniques, with the exception of certain specialist applications.
Further ahead, one of the biggest developments is the maturation of digital PCR technology. Digital PCR has become much easier to deploy and is a technique we are definitely keeping an eye on. However there are some clear limitations, not least of all the high cost of the instruments and the complexity of the process. For this reason instruments like the Eco 48 still have clear advantages; qPCR is not only fast and robust but the software makes handling the reaction very straightforward. Moreover as most qPCR assays involve looking at the relative expression of one message over another it simply is not necessary to have the absolute precision in the quantity of target copies generated by digital PCR. I cannot see digital PCR being taken out into the field to monitor pathogens, for instance, while there are already hand-held battery driven qPCR devices available to perform such experiments. Because of its ubiquity and huge adaptability in terms of instrumentation, reagents and application it is safe to say we will still be using qPCR in ten years’ time.
The technology behind the research
With sensitivity of detection down to a single copy and the world’s most thermally accurate block, the Eco 48 from PCRmax enables researchers to surpass all required MIQE guidelines and accelerate run time, with no compromise to efficiency.
This is achieved through a number of innovations, including novel block technology which delivers complete heating uniformity and rapid cycles in under 40 minutes, with the sensitivity to deliver ±0.1oC uniformity across the whole block instantly after every temperature change (Figure 1).

Meanwhile adaptive LED control a the high-performance optical system removes destructive variables from the fluorescence detection process, such as unknown saturation or light bleaching through to neighbouring cells (Figure 2).

Combined with intutive software, designed with ease of use in mind, the Eco 48 makes it faster and easier to develop rigorously validated assays that deliver complete confidence in results, improving the anaytical productivity for practically all qPCR application.
PCRmax is a Bibby Scientific company specialising in providing real time instrumentation, consumables and reagents for the life science laboratory. Find out more about PCRmax and the Eco 48 at www.pcrmax.com
For more information on Professor Bustin’s research contact enquiry@pcrmax.com
References
[1] Bustin S.A. et.al; The MIQE guidelines: minimum information for publication of quantitative real-time PCR experiments Clin Chem. 2009 Apr; 55(4):611-22

