Structure of Cold Virus Solved

Complete the form below to unlock access to ALL audio articles.
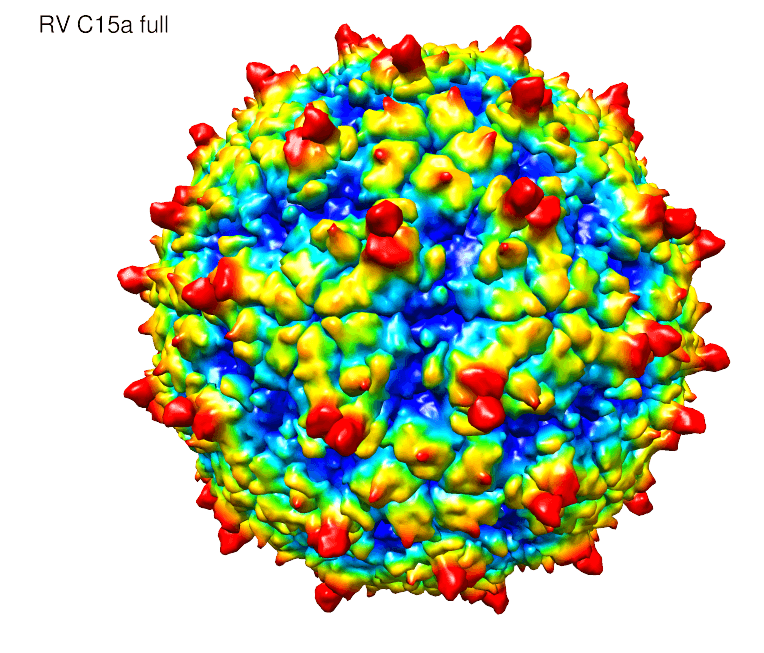
The atomic structure of an elusive cold virus linked to severe asthma and respiratory infections in children has been solved by a team of researchers at the University of Wisconsin–Madison and Purdue University.
The findings are published this week in the Proceedings of the National Academy of Sciences (PNAS) and provide the foundation for future antiviral drug and vaccine development against the virus, rhinovirus C.
“Rhinovirus C has been the ‘missing link’ in explaining illness caused by the common cold,” says Michael Rossmann, Hanley Distinguished Professor of Biological Science at Purdue and co-lead of the study with UW–Madison’s Ann Palmenberg, professor of biochemistry and with the Institute for Molecular Virology. “This is sure to spark major new efforts toward the development of vaccines and antiviral drugs, particularly for the prevention of asthma.”
In fact, multiple pharmaceutical companies have been waiting for the data, Palmenberg says.
Rhinovirus C is resistant to current antiviral drugs and no vaccines exist. It was discovered just 10 years ago and health experts believe it is responsible for 50 to 85 percent of all childhood hospitalizations for asthma.
Asthma affects an estimated 24 million people in the U.S. and asthmatic children infected with rhinovirus C have been shown in previous studies to have more severe asthma symptoms.
Rhinovirus C infections are also a major contributor to poor outcomes for people with chronic lung conditions like cystic fibrosis, and early exposure to the virus can cause lifelong lung scarring leading to persistent respiratory difficulties later, says Palmenberg.
Last fall, using a rhinovirus C preparation grown in the Palmenberg lab — currently one of the few places in the world capable of culturing the virus — and an imaging technology new to Purdue, Rossmann’s graduate student, Yue Liu, first author of the study, was able to map the full atomic structure of a rhinovirus C particle.
Discerning the structure of microscopic biological agents like viruses allows them to be studied and understood in ways not possible by other means. For instance, Liu uncovered physical features of rhinovirus C that help explain why it is resistant to standard antiviral treatments, how it interacts with cells to cause infection, and why it behaves as a different disease from other members of the enterovirus family.
“We found some interesting things,” says Palmenberg. “Unlike normal rhinoviruses, this one has spikes on the surface of the particles. We had not anticipated that.”
This and other virus surface changes help explain why clinical rhinovirus drug trials, which rely on different virus surfaces, have generally failed when tested against rhinovirus C isolates over the last four to five years, says Palmenberg.
However, the spikes present new binding sites for the human immune response to infection and as such, may present promising new targets for the development of different types of drugs. Or, they may contain structure elements that could be leveraged for effective vaccines.
“Another new finding is that 30 percent of the virus particles were empty and do not contain genetic material,” Palmenberg says.
The “empty” particles are not capable of causing infection, but like tiny molecular mimics, they present virus-specific information to the immune system.
“They could be immune decoys,” says Palmenberg, luring the body to fight off noninfectious viral particles, leaving infectious virus free to do its dirty work. “No other rhinoviruses seem to do this naturally.”
They could potentially be used for vaccines.
The virus was among the first samples analyzed with a new cryo-electron microscopy system at Purdue, which was designed to operate at vastly higher resolution than previous instruments. The system allows for multiple imaging of single virus particles, rather than relying on a composite of images from thousands of individual viruses. Therefore, the system is faster, more accurate and gives higher resolution structures from less virus material.
While the rhinovirus C structure determination was certainly accelerated by this new technology, it was also dependent upon Rossmann’s many years of expertise, Liu’s efforts, and years of work by Palmenberg’s experienced team in order to get the virus to grow under laboratory conditions.
“If you can’t grow it, you can’t do experiments,” says Palmenberg. “That was the linchpin for this structure … it took us two-and-a-half years of work to figure out how to put sufficient virus into a tube so we could give it to Purdue.”
That process has now been licensed by the Wisconsin Alumni Research Foundation, a private nonprofit that manages intellectual property at UW–Madison.
While learning how to culture rhinovirus C, Palmenberg’s research team discovered the virus only infects certain lung cells that express a particular genetic form of a cell-surface molecule called cadherin-related protein 3 (CDHR3). The team published those results last year in PNAS.
When a rhinovirus C particle encounters a cell bearing the right CDHR3 protein, it can lock onto and gain entry into the cell. During the ensuing infection, it makes many copies of its genetic material, using the cell’s machinery, and this reassembles into many more virus particles that are then released to infect more cells.
When CDHR3 is expressed on cells deep in the lungs, the virus infections can wreak havoc, says Rossmann. “It creates the perfect storm for the severest type of asthma-inducing, lung-scarring events.”
But “knowing how to grow it doesn’t tell you what it looks like,” says Palmenberg. Multiple types of rhinovirus C exist and they are part of a larger group of illness-causing viruses, including cold viruses, rhinovirus A and B. “We knew it would be different from rhinovirus A and B: It didn’t respond to drugs, it used a different receptor, it had different biological properties. So, what was going on?”
With the new structure data, the team is much closer to finding answers. In fact, mapping the rhinovirus C particles also showed where on these surfaces the virus likely makes contact with the CDHR3 molecules, presenting yet another opportunity to design drugs which may prevent infection.
Palmenberg and Rossmann have collaborated for 34 years, ever since her former mentor, UW–Madison professor emeritus Roland Rueckert, and Rossmann collaborated to become the first to determine the structure of rhinovirus B.
Once Palmenberg worked out how to grow rhinovirus C, she, too, partnered with Rossmann. Her research technician and co-author of the paper, Marchel Hill, delivered a crude test sample of the virus to Purdue on a Friday morning last fall. Liu performed some of the imaging that same day and a preliminary structure were mostly ready by Saturday.
The study was also co-authored by UW–Madison’s Kelly Watters; Purdue’s Thomas Klose, Zhenguo Chen and Wen Jiang; and Yury Bochkov of the UW–Madison School of Medicine and Public Health. It was funded by the National Institutes of Health.

