Single Cell Isolation Trends: Technologies, Limitations & Applications

Complete the form below to unlock access to ALL audio articles.
Introduction
Historically cell populations were considered homogeneous and conventional cell-based assays utilized the average responses from a population of many thousands of cells. The results obtained from such analyses represent an amalgam of the biological status of each cell within the population analysed without considering individual cell phenotypes. To better understand the variations from cell to cell, scientists need to study single cells. Recent advances of single cell analysis techniques have facilitated the ability to discern biological insights within individual cells and provided the means to revealing previously hidden relationships between individual cells within a population or to detect subpopulations. Minority, rare cell events, and small changes between individual cells may hold the key to answering hitherto unresolved questions in cancer, stem cell biology, immunology, developmental biology and neurology, and facilitate therapeutic decision making in precision medicine.
Multi-application µEncapsulator System - simple, quick and reliable encapsulation of single cells

The µEncapsulator System enables simple, quick and reliable encapsulation of single cells, DNA and/or functionalized beads in high precision, monodisperse picoliter droplets and can encapsulate 300,000 cells in 15 minutes. The system is Ideal for a wide range of applications including profiling natively paired TCRs, isolating antibody coding sequences, encapsulating cells in hydrogels or encapsulating expression libraries for FACS sorting.
“We have developed a flexible high throughput single cell product which provides customers with simple single cell encapsulation for a wide range of applications enabling high throughput single cell profiling."
Mike Hawes, CEO, Dolomite Bio
Prior to initiating single cell analysis, scientists need to separate and isolate single cells. Cell isolation technologies are characterized by 3 performance criteria: efficiency or throughput (i.e. the number of single cells that can be isolated in a certain time); purity (i.e. the % of isolated cells that have the desired phenotype); and recovery (i.e. the % of target cells isolated relative to the total number of target cells available in the starting sample). Current cell isolation techniques broadly fall into 2 categories: 1) isolation based on physical properties like size, density, electric changes, and deformability, with methods including density gradient centrifugation, membrane filtration, microchip-based capture platforms, laser microdissection, manual cell picking/micromanipulation and non-contact dispensing; and 2) isolation based on cellular biological characteristics and protein expression properties. These comprise affinity methods that typically involve labelling, such as affinity matrices, fluorescence-activated cell sorting and magnetic-activated cell sorting, many are based on microfluidic platforms. The most wanted single cells are however those that are isolated unlabelled and retain full viability. New miniaturized all-in-one single cell platforms that enable both the isolation and analysis are proving of increasing interest to researchers, particularly in the area of single cell genomics.
In December 2016 HTStec undertook a market survey on single cell isolation mainly among research labs in academia, pharma and biotech. The objectives were to understand current interest, progress made in implementation and requirements for single cell isolation. This article contains ‘selected findings’ from the HTStec market report ‘Single Cell Isolation Trends 2016’. It is intended to provide the reader with a brief insight into recent market trends. It covers only 11 out of the 31 original questions detailed in the full report. The full published report should be consulted to view the entire dataset, details of the breakdown of the response to each question, its segmentation and the estimates for the future. Please contact info@htstec.com for more information about HTStec reports.
Sources used to isolate single cells:
The sources most used by survey respondents to isolate single cells are presented in Figure 1. This showed that suspension cell cultures were most used (45% utilizing). This was followed by adherent tissue cultures (39%), clinical samples - fresh tissues (29%), and then clinical samples - fresh biofluids (27%). Other sources were used by less than 20% of survey respondents.
 Figure 1. Sources Used To Isolate Single Cells
Figure 1. Sources Used To Isolate Single CellsCell types used for single cell technology applications:
The main cell types used for single cell applications are reported in Figure 2. The cell type from which respondents most want to isolate single cells was primary cells. This was followed by stem cells or stem cell-derived cells (37%), and then tumor cell lines (e.g. Hela, U937 etc.) (33%). Other cell types were used by less than 20% of survey respondents. There was limited interest in non mammalian cell types.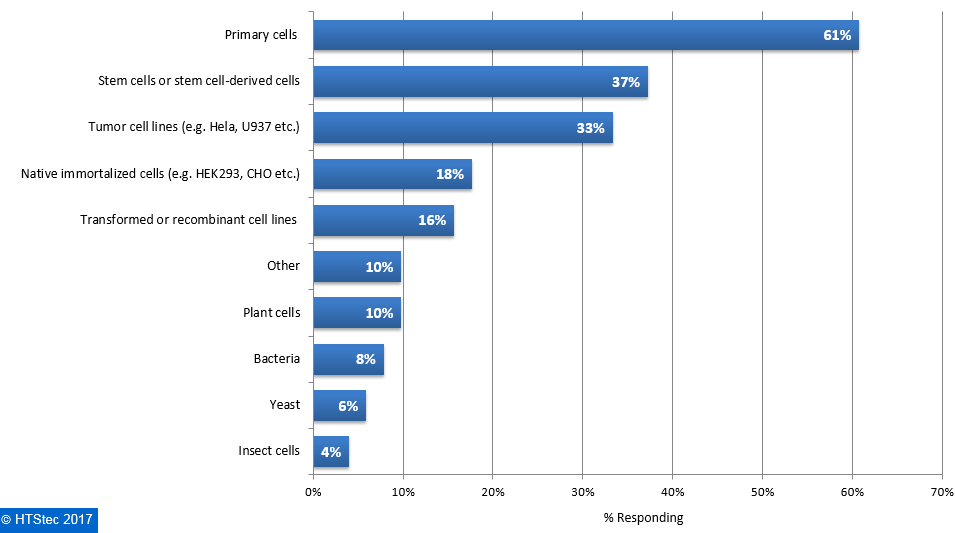 Figure 2. Cell Types Used For Single Cell Technology Applications
Figure 2. Cell Types Used For Single Cell Technology Applications
The starting population from which single cells are isolated:
The starting population from which survey respondents most want to isolate single cells are given in Figure 3. This showed that an abundant representative sample, just need to isolate individual cells was most preferred (48% preferring). This was followed by an abundant sample looking for rare events or a precious sample in very low quantities (both with 26% preferring). 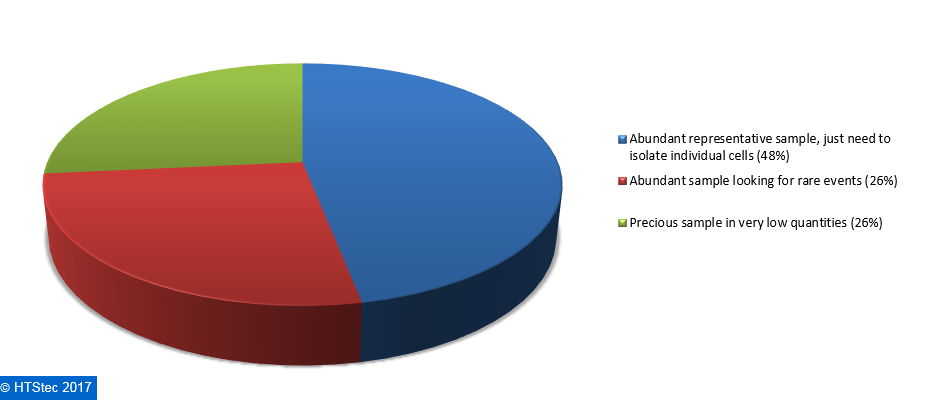 Figure 3. The Starting Population (Sample) From Which Respondents Isolate Single Cells
Figure 3. The Starting Population (Sample) From Which Respondents Isolate Single Cells
The main technologies used to isolate single cells:
The main technologies currently used to isolate single cells are presented in Figure 4. This showed that FACS/flow cytometry was currently the most used technology (41% using). This was followed by microfluidic/lab-on-a-chip (29% using), manual cell picking/micro-manipulation (12% using), other technologies (not specified) (10% using), laser microdissection (4% using), and then random seeding or liquid dilution into microplates and non-contact dispensing methods (e.g. inkjet, solenoid valves or acoustic) (both with 2% using).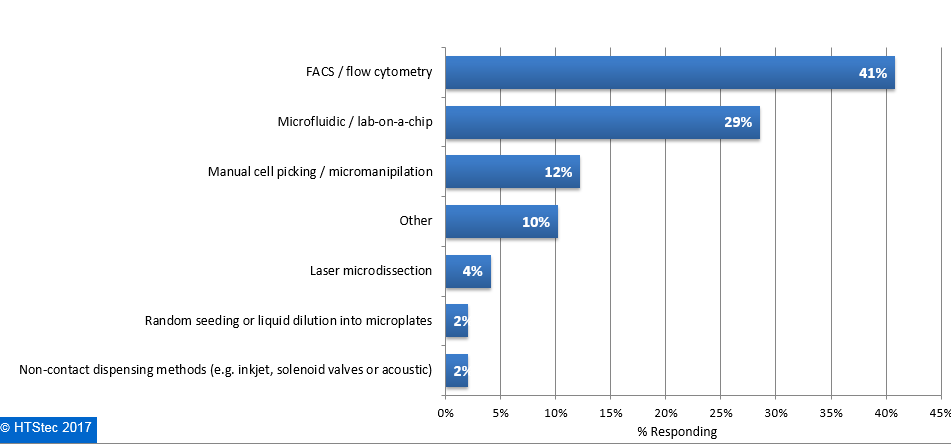 Figure 4. Main Technology Currently Used To Isolate Single Cells
Figure 4. Main Technology Currently Used To Isolate Single Cells
The category of cell purity of greatest relevance:
The category of cell purity of greatest relevance to survey respondent’s research on isolated single cells is reported in Figure 5. This showed that cells with desired phenotype were ranked most relevant. This was followed in relevance by cells that are singlet (i.e. not clumped), cells that are fully viable and then cells free from attached debris. Least relevant was cells at cell cycle specific stage of development.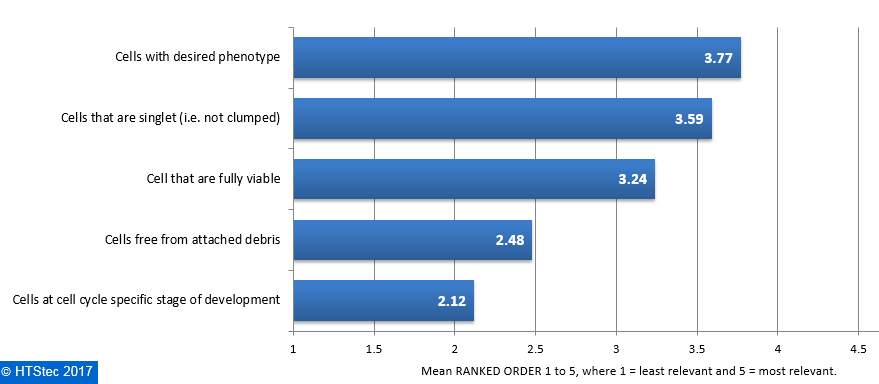 Figure 5. Category Of Cell Purity Of Greatest Relevance To Respondent's Research On Isolated Single Cells
Figure 5. Category Of Cell Purity Of Greatest Relevance To Respondent's Research On Isolated Single Cells
The importance of isolating individual cells:
The importance of isolating individual cells for downstream experiments are given in Figure 6. This revealed it was of critical importance to most survey respondents research i.e. they absolutely need single cells (47% selecting). This was followed by desirable i.e. they can use a couple (1-5) of cells as long as they have the same phenotype (37% selecting), and then useful i.e. they will pool the selected cells together to minimize the need for amplification (14% selecting). The remaining respondents (2% selecting) indicated it was not important i.e. they are OK with a small number of cells regardless of their phenotype.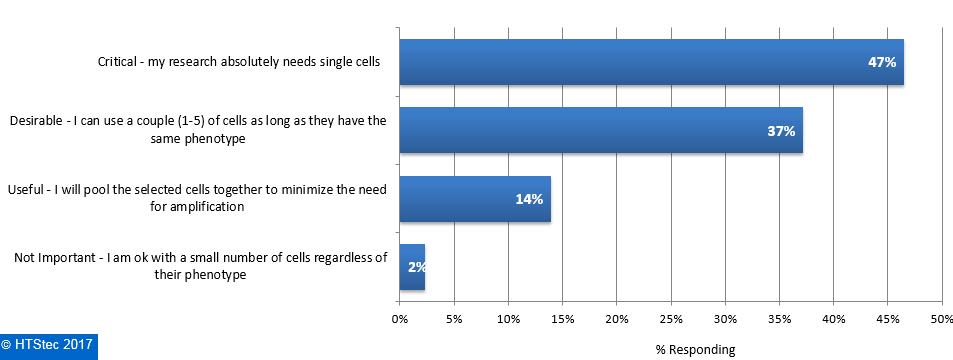 Figure 6. Importance Of Isolating Individual Cells For Downstream Experiments
Figure 6. Importance Of Isolating Individual Cells For Downstream Experiments
The importance of isolating viable single cells:
The importance of isolating viable single cells to survey respondents planned research is detailed in Figure 7. This showed that the majority (51%) reported that it was critical to maintain the viability of isolated cells, 40% said it was nice to have, and only 9% reported that live cells are not needed for their research. These findings were supported by the finding that at least 42% of respondents’ research involved further maintenance/culture of living cells e.g. to collect secreted proteins. Figure 7. The Importance Of Isolating Viable Single Cells
Figure 7. The Importance Of Isolating Viable Single Cells
The main downstream analysis methods to be used:
The main downstream analysis methods used or planned to be used with isolated single cells are presented in Figure 8. This showed that the main downstream application used was RT-qPCR (51% using). This was followed by cellular phenotype determination and counting (49% using), NGS: RNA sequencing (33% using), protein analysis with labeled antibodies (26% using), protein analysis with labeled antibodies (26% using), NGS: whole genome sequencing (23% using), genotyping (16% using), and then other (14% using).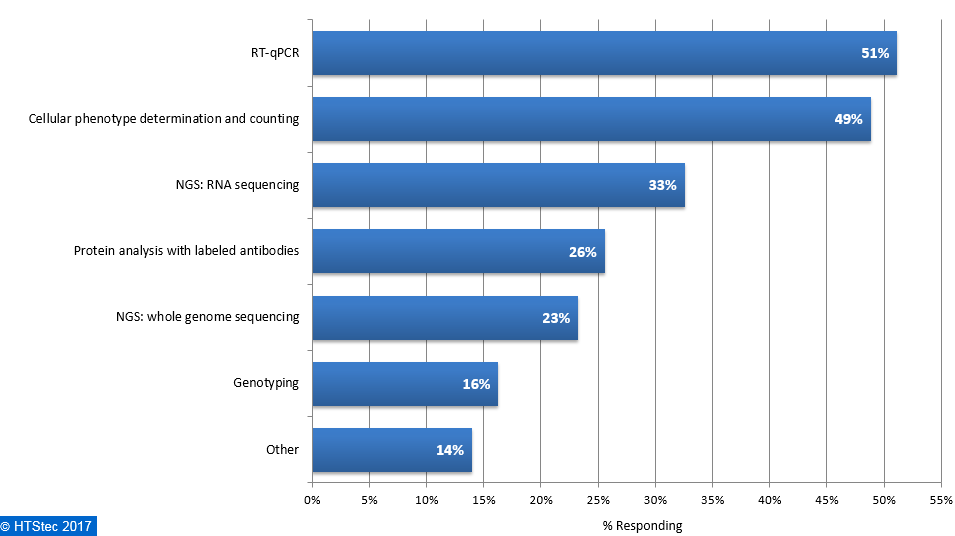 Figure 8. Downstream Analysis Methods Used Or Planned To Be Used With Isolated Single Cells
Figure 8. Downstream Analysis Methods Used Or Planned To Be Used With Isolated Single Cells
Vendor awareness in single cell isolation technologies:
The supplier/vendor of single cell isolation technologies that first comes into the mind of survey respondents is presented in Figure 9. This showed that BD Biosciences and Fluidigm/Bio-Rad were the most recognized vendors of single cell technologies (each with 16% selecting). This was followed by Miltenyi Biotech (6%); 10X Genomics (4%); Beckman Coulter (4%); Stemcell Technologies (4%); Thermo Fisher Scientific (3%); Zeiss (3%); GE Healthcare (3%); and then all other suppliers (each with less than 1% share) (41%). Please note this should NOT be regarded as a true market share projection. It is based on the supplier of single cell isolation technologies that first comes into the mind of survey respondents, which may not be the same as those who they purchase most consumables or instruments from. 
Figure 9. Supplier/Vendor Of Single Cell Isolation Technologies That First Comes To Mind
Application most likely to be impacted by single cell isolation technologies:
The downstream application most likely to be impacted by single cell isolation technologies are given in Figure 10. This showed that survey respondents equally selected biomarker discovery, validation and/or screening and gene expression profiling as the most impacted applications (both with 11% selecting). They were followed by single cell heterogeneity (10% selecting) and then genome and transcriptome analysis, RNA/DNA sequencing, molecular characterization of circulating tumor cells and single cell immunology (all with 7% selecting) and then cell line development (6% selecting). All other downstream applications were selected by 5% or less of respondents.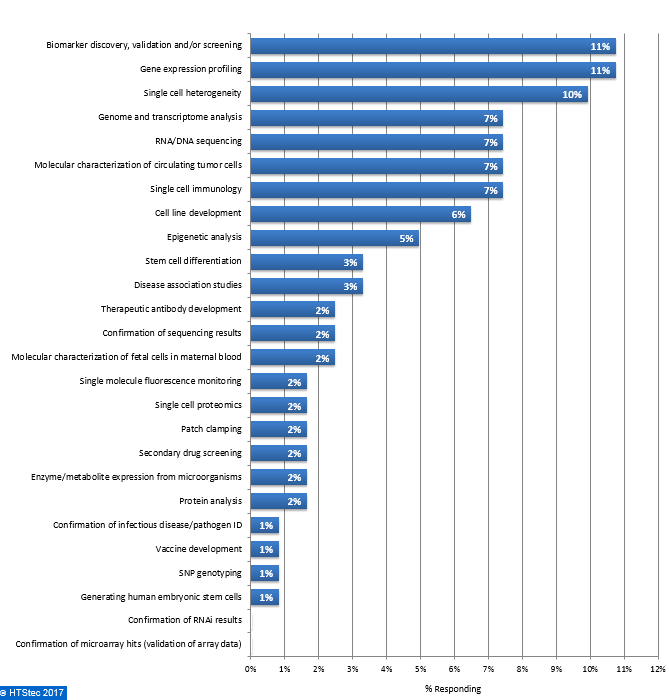 Figure 10. Downstream Application Most Likely To Be Impacted By Single Cell Isolation Technologies
Figure 10. Downstream Application Most Likely To Be Impacted By Single Cell Isolation Technologies
Biggest limitations of available technologies:
The biggest limitations of currently available single cell isolation technologies were rated in Figure 11. This showed that poor efficiency or low yield/recovery was rated the most limiting. This was followed in the ratings by high cost of equipment, and then degradation of cell viability/functionality in the separation process, poor reliability, and low throughput (in terms of single cells isolated per second). All other factors were rated moderately limiting, suggesting they all represent significant problems in single cell isolation. Figure 11. Main Limitations Of Currently Available Single Cell Isolation Technologies
Figure 11. Main Limitations Of Currently Available Single Cell Isolation Technologies
Conclusions:
The selected findings reported above revealed suspension cell cultures were still most used to isolate single cells. The main cell types used for single cell applications were primary cells. The preferred starting population was an abundant representative sample i.e. they need to isolate individual cells. The main technology currently used to isolate single cells was FACS/flow cytometry. The category of cell purity of greatest relevance to survey respondent’s research were cells with a desired phenotype. The importance of isolating individual cells for downstream experiments was of critical importance to most survey respondents research i.e. they absolutely need single cells. The majority of respondents reported that it was critical to maintain the viability of isolated single cells. The main downstream analysis method used or planned to be used with isolated single cells was RT-qPCR. The supplier/vendor of single cell isolation technologies that first comes into the mind of survey respondents was BD Biosciences and Fluidigm/Bio-Rad. The downstream applications most likely to be impacted by single cell isolation technologies were biomarker discovery, validation and/or screening and gene expression profiling. Poor efficiency or low yield/recovery was rated the biggest limitation of currently available single cell isolation technologies.
Although much progress has been made during the recent years in single cell isolation there are still major challenges with isolation methodologies. In particular the isolation efficiency, the high cost of equipment and degradation of cell viability/functionality in the separation process are all reported as major limitations and areas where respondents are seeking improvement. Several new and emerging technologies now available to isolate viable single cells address some of these issues:
Menarini Silicon Biosystems DEPArray™ technology which is based on the ability of a non-uniform electric field to exert forces on neutral, polarizable particles, such as cells, that are suspended in a liquid. This electrokinetic principle, called dielectrophoresis (DEP), can be used to trap cells in DEP “cages” by creating an electric field above a subset of electrodes in an array that is in counter phase with the electric field of adjacent electrodes. When a DEP cage is moved by a change in the electric field pattern, the trapped cell moves with it.
Cytena Single Cell Printer enables fully automated isolation of single cells into standard microwell plate formats. The instrument uses an inkjet-like principle featuring a disposable, one-way printing cartridge. The cell sample is pipetted into the cartridge and an external actuator is used to eject droplets out of it. The integrated optical sensor allows for determination of cell number in each droplet. A fast shutter mechanism sorts the droplets containing exactly one single-cell into the substrate. Unwanted droplets are deflected into waste.
QIAGEN QIAscout core technology is the QIAscout array, which is composed of 12,000 individual microrafts. Each microraft is 200 µm in length and 200 µm in width and is securely held within a micron-scale well of the QIAscout array. Within the QIAscout array, the microrafts serve as releasable culture sites for individual cells or colonies. QIAscout arrays are mounted on inverted microscopes and can be imaged by brightfield, fluorescence and confocal microscopy. A motorized release device is mounted onto a 4x, 5x or 10x objective of common inverted lab microscopes to allow for selection and picking of individual microrafts. To isolate target cells, simply press the button on the controller which makes the release needle pierce through the lower surface of the QIAscout array and dislodge an individual microraft with your cell of interest attached. The released microraft is then recovered using a magnetic wand and transported to a secondary vessel (e.g. reaction tube or microplate well for downstream analysis).
Sphere Fluidics Picodroplet Single Cell Encapsulation System. This semi-automated system encapsulates single cells or biomolecules into picodroplets, ready for downstream screening and analysis. It does this at a rapid rate of up to 70,000 picodroplets per second, and the flow rate can be highly controlled. The encapsulation system doesn’t affect cell viability, while the wide range of picodroplet sizes and volumes gives it flexibility for use with various cell types – large or small. These picodroplets can be stabilised using novel surfactants and cells can be grown in them, and even incubated or stored for many days.
Over the coming years we can expect single cell techniques will become a powerful tool in unravelling longstanding questions in our understanding and treatment of human disease.
DISCLAIMER: HTStec Limited has exercised due care in compiling and preparing these Selected Findings from its Report, which is based on information submitted by individuals in respondent companies. HTStec Limited has NOT verified the accuracy of this information, nor has it is established respondent’s authority to disclose information to HTStec Limited. HTStec Limited expressly disclaims any and all warranties concerning these Selected Findings including any warranties of merchantability and/or fitness for any particular purpose, and warranties of performance, and any warranty that might otherwise arise from course of dealing or usage of trade. No warranty is either expressed or implied with respect to the use of these Selected Findings. Under no circumstances shall HTStec Limited be liable for incidental, special, indirect, direct or consequential damages or loss of profits, interruption of business, or related expenses that may arise from use of these Selected Findings, including but not limited to those resulting from inaccuracy of the data therein.

