Study Identifies Oncoprotein’s Cryptic Vulnerabilities for Anti-Cancer Drug Development
Complete the form below to unlock access to ALL audio articles.
RET is a receptor protein with tyrosine kinase activity that can transfer a phosphate group from ATP to other molecules modifying their shape and function in signalling pathways that are essential to organogenesis and tissue maintenance. Gain-of-function genetic alterations in RET, e.g point mutations or chromosomal rearrangements that produce oncogenic fusions, are implicated in some types of cancer, especially those of thyroid, and less frequently, lung and breast cancers, among others. Current anticancer therapies agaisnt RET-driven tumours are based on ATP-competitive inhibitors of the RET catalytic activity. Second generation inhibitors, which include LOXO-292 (selpercatinib) and BLU-667 (pralsetinib), have been approved by the FDA and result in remarkable clinical responses in cancer patients.
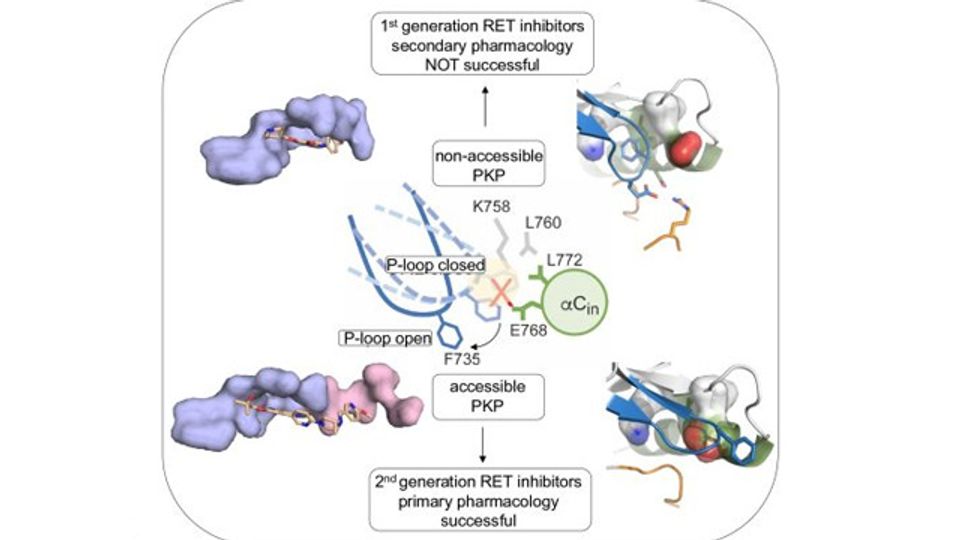
The Kinases, Protein Phosphorylation and Cancer Group at the Spanish National Cancer Research Centre (CNIO), led by Ivan Plaza Menacho, has successfully identified structural and dynamic features exploited by these inhibitors, and in particular a cryptic pocket within the active site that conferred high specificity to these compounds. “The crystal structure of the RET catalytic domain in complex with these inhibitors were already solved, but the cryptic pockets in the active site were not identified, since they are not always accessible due to the dynamic and conformational changes that the crystal structure does not capture,” Plaza-Menacho remarks.
The Kinases, Protein Phosphorylation and Cancer Group is focused on the detailed structural and functional characterization of a family of proteins called kinases that are implicated in cancer. This information can be translated into the design of better compounds and inhibitors that will result in improved therapies. “Second-generation RET inhibitors, which have been recently approved by the FDA to treat RET-driven thyroid and lung cancers, bind to the active site in an unusual way compared with prototypical RET inhibitors. However, we did not fully understand why they were so effective,” Plaza-Menacho adds.
Currently, the group led by Plaza-Menacho is working in close collaboration with the Experimental Therapeutics Programme at CNIO on the identification, design and development of new compounds targeting the identified cryptic pocket and other allosteric sites. We expect some of these hits will lead to drugs that can be potentially used in precision and personalised therapies to treat RET-driven cancers in the future, said Plaza-Menacho. Refractory mutations conferring resistance to these inhibitors have already been described. The authors in the study also provide some guidelines for the design of new drugs that can overcome the effects of such refractory mutations.
Reference: Shehata MA, Contreras J, Martín-Hurtado A, et al. Structural and dynamic determinants for highly selective RET kinase inhibition reveal cryptic druggability. J. Adv. Res. 2022. doi: 10.1016/j.jare.2022.05.004
This article has been republished from the following materials. Note: material may have been edited for length and content. For further information, please contact the cited source.

