Rare Form: Novel Structures Built from DNA Emerge

Complete the form below to unlock access to ALL audio articles.
The technique, known as DNA origami, promises to bring futuristic microelectronics and biomedical innovations to market.
Hao Yan, a researcher at Arizona State University’s Biodesign Institute, has worked for many years to refine the technique. His aim is to compose new sets of design rules, vastly expanding the range of nanoscale architectures generated by the method. In new research, a variety of innovative nanoforms are described, each displaying unprecedented design control.
Yan is the Milton D. Glick Distinguished Chair of Chemistry and Biochemistry and directs Biodesign’s Center for Molecular Design and Biomimetics.
In the current study, complex nano-forms displaying arbitrary wireframe architectures have been created, using a new set of design rules.
“Earlier design methods used strategies including parallel arrangement of DNA helices to approximate arbitrary shapes, but precise fine-tuning of DNA wireframe architectures that connect vertices in 3D space has required a new approach,” Yan says.
Yan has long been fascinated with Nature’s seemingly boundless capacity for design innovation. The new study describes wireframe structures of high complexity and programmability, fabricated through the precise control of branching and curvature, using novel organizational principles for the designs. (Wireframes are skeletal three-dimensional models represented purely through lines and vertices.)
The resulting nanoforms include symmetrical lattice arrays, quasicrystalline structures, curvilinear arrays, and a simple wire art sketch in the 100-nm scale, as well as 3-D objects including a snub cube with 60 edges and 24 vertices and a reconfigurable Archimedean solid that can be controlled to make the unfolding and refolding transitions between 3-D and 2-D.
In previous investigations, the Yan group created subtle architectural forms at an astonishingly minute scale, some measuring only tens of nanometers across – roughly the diameter of a virus particle. These nano-objects includespheres, spirals, flasks, Möbius forms, and even an autonomous spider-like robot capable of following a prepared DNA track.
The technique of DNA origami capitalizes on the simple base-pairing properties of DNA, a molecule built from the four nucleotides Adenine (A), Thymine (T) Cytosine (C) and (Guanine). The rules of the game are simple: A’s always pair with T’s and C’s with G’s. Using this abbreviated vocabulary, the myriad body plans of all living organisms are constructed; though duplicating even Nature’s simpler designs has required great ingenuity.
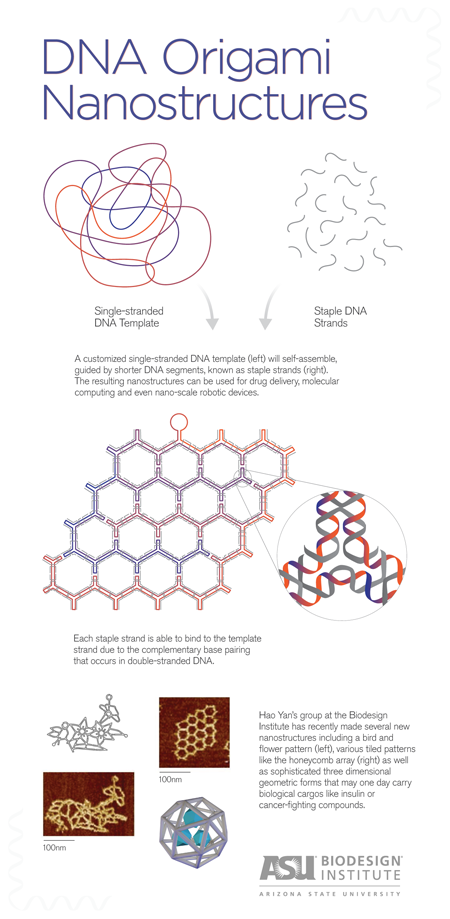
The basic idea of DNA origami is to use a length of single-stranded DNA as a scaffold for the desired shape. Base-pairing of complementary nucleotides causes the form to fold and self-assemble. The process is guided by the addition of shorter “staple strands,” which act to help fold the scaffold and to hold the resulting structure together. Various imaging technologies are used to observe the tiny structures, including fluorescence-, electron- and atomic force microscopy.
Although DNA origami originally produced nanoarchitectures of purely aesthetic interest, refinements of the technique have opened the door to a range of exciting applications including molecular cages for the encapsulation of molecules,enzyme immobilization and catalysis, chemical and biological sensing tools, drug delivery mechanisms, and molecular computing devices.
The technique described in the new study takes this approach a step further, allowing researchers to overcome local symmetry restrictions, creating wireframe architectures with higher order arbitrariness and complexity. Here, each line segment and vertex is individually designed and controlled. The number of arms emanating from each vertex may be varied from 2 to 10 and the precise angles between adjacent arms can be modified.
In the current study, the method was first applied to symmetrical, regularly repeating polygonal designs, including hexagonal, square and triangular tiling geometries. Such common designs are known as tessellation patterns. A clever strategy involving a series of bridges and loops was used to properly route the scaffold strand, allowing it to pass through the entire structure, touching all lines of the wireframe once and only once. Staple strands were then applied to complete the designs.
In subsequent stages, the researchers created more complex wireframe structures, without the local translational symmetry found in the tessellation patterns. Three such patterns were made, including a star shape, a five-fold Penrose tile and an eight-fold quasicrystalline pattern. (Quasicrystals are structures that are highly ordered but non-periodic. Such patterns can continuously fill available space, but are not translationally symmetric.)
Loop structures inserted into staple strands and unpaired nucleotides at the vertex points of the scaffold strands were also used, allowing researchers to perform precision modifications to the angles of junction arms.
The new design rules were next tested with the assembly of increasingly complex nanostructures, involving vertices ranging from two to 10 arms, with many different angles and curvatures involved, including a complex pattern of birds and flowers.
The accuracy of the design was subsequently confirmed by AFM imaging, proving that the method could successfully yield highly sophisticated wireframe DNA nanostructures. The method was then adapted to produce a number of 3-D structures as well, including a cuboctahedron, and another Archimedian solid known as a snub cube – a structure with 60 edges, 24 vertices and 38 faces, including six squares and 32 equilateral triangles.

