Gram Positive vs Gram Negative
One common way to separate bacteria is based on the structure of bacterial cell walls. Let's take a closer look.

Complete the form below to unlock access to ALL audio articles.
Credit: Technology Networks/YouTube
Being able to differentiate bacterial species is important for a host of reasons, from diagnosing infection or checking food safety, to identifying which species it is that gives a cheese it’s fantastic character. Bacterial species, and even specific strains can be differentiated using a number of molecular techniques such as PCR, quantitative PCR, genome sequencing and mass spectrometry. But even without getting into the molecular nitty gritty, there are phenotypic differences between groups of bacteria that can be used to differentiate them. This includes characteristics like their shape (bacilli vs cocci for example), growth in particular nutrients and preference for high or low oxygen environments. Depending on the characteristic being studied, bacterial species may be broken down into broad groups, but taken together this information can narrow the possible identities greatly. One such useful classification – if a bacterium is Gram positive or Gram negative - is based on the structure of bacterial cell walls.
Contents
Gram positive vs gram negative
Difference in structure of Gram positive vs Gram negative bacteria
Gram positive vs Gram negative stain
- Gram stain procedure - Preparing a sample
- Gram stain procedure - Gram staining a sample
Gram positive vs gram negative
Gram positive bacteria are surrounded by a single thick peptidoglycan cell wall and are therefore termed monoderms. Gram negative bacteria have a much thinner peptidoglycan cell wall, but in addition they have an outer membrane containing lipopolysaccharides surrounding the cell and are consequently termed diderms.
Gram positive bacteria | Gram negative bacteria |
| Distinctive purple appearance after gram staining | Pale reddish color after gram staining |
| Bacteria include all staphylococci, all streptococci and some listeria species | Bacteria include enterobacter species, salmonella species and pseudomonas species |
| Thick peptidoglycan layer | Thin peptidoglycan layer |
| No outer lipid membrane | Outer lipid membrane present |
| No O-specific side chains present | O-specific side chains present |
| Teichoic and lipoteichoic acids present | Teichoic and lipoteichoic acids not present |
Difference in structure of Gram positive vs Gram negative bacteria
The diagram below illustrates the differences in the structure of Gram positive and Gram negative bacteria. The two key features that lead to the differing visualization properties of Gram positive and Gram negative species are the thickness of the peptidoglycan layer and presence or absence of the outer lipid membrane. This is because the wall structure affects the cell’s ability to retain the crystal violet stain used in the Gram staining procedure which can then be visualized under a light microscope.
 Credit: Technology Networks
Credit: Technology Networks
Gram positive bacteria have a thick peptidoglycan layer and no outer lipid membrane whilst Gram negative bacteria have a thin peptidoglycan layer and have an outer lipid membrane.
As Gram positive bacteria lack an outer lipid membrane, when correctly referring to their structure rather than staining properties, are termed monoderms. The outer lipid membrane possessed by Gram negative bacteria means that, when referring to their physical structure, they are termed diderms.
The Gram staining technique was developed in 1884 by Danish bacteriologist Hans Christian Gram.1 Whilst a Gram stain will not tell you the specific species you are looking at, it can be a quick way to narrow down greatly the list of potential candidates and direct follow-up testing where necessary.
Gram positive vs Gram negative stain
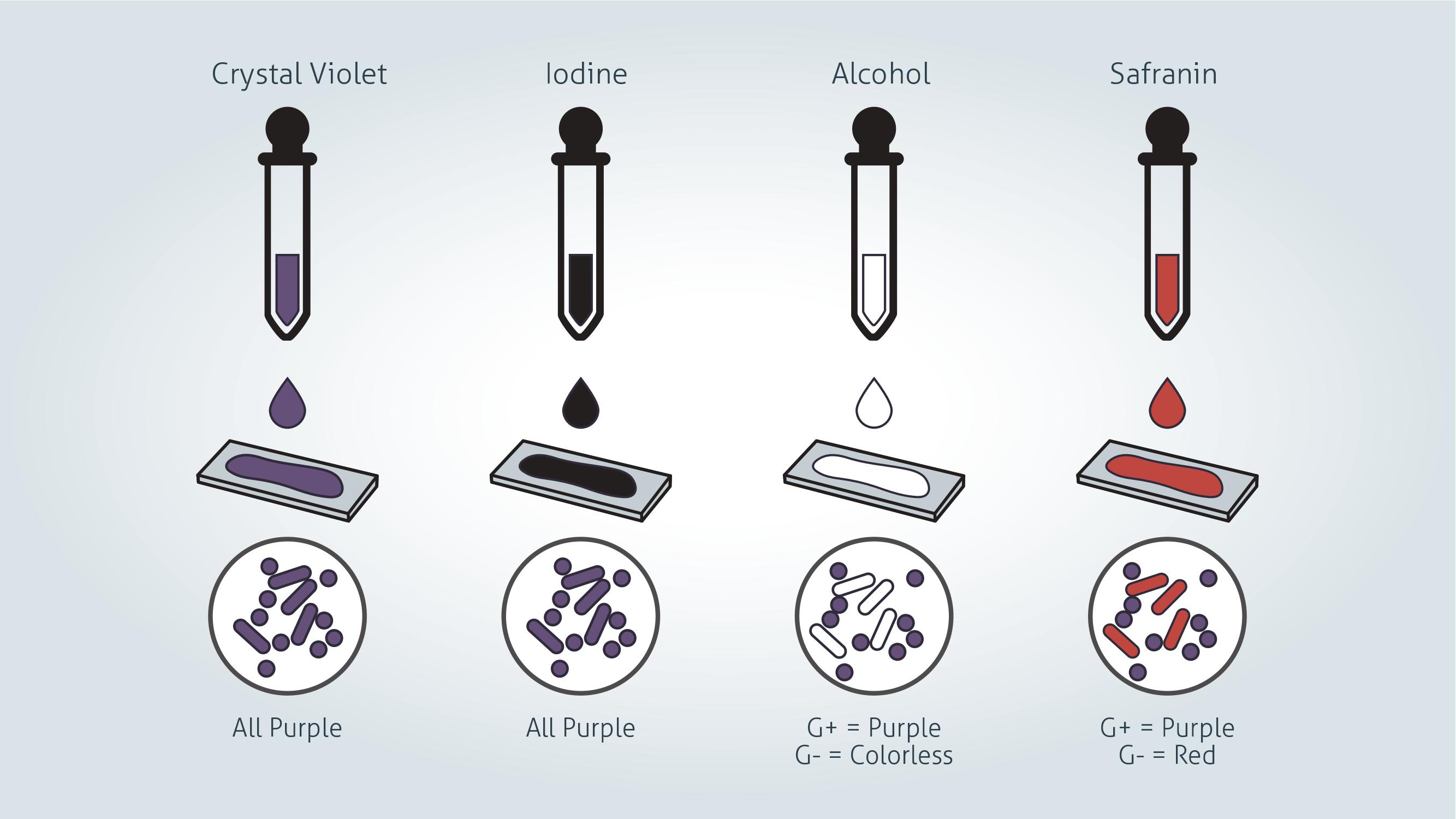 Credit: Technology Networks
Credit: Technology NetworksGram stain procedure - Preparing a sample
1. Label a clean glass microscope slide with your sample identification. Ensure you use a pencil as many inks are removed by the reagents used in the staining procedure.
2. If preparing your slide from a liquid bacterial culture:
Dab a small drop culture onto the slide using a sterile loop. Gently smear the droplet in a circular motion into an area of approximately 1 cm diameter. For very dense cultures it may be necessary to pre-dilute your culture to ensure individual bacterial cells can be seen under a microscope following staining.
If the source material is from a bacterial plate:
Resuspend a loop of colony material in sterile phosphate buffered saline (PBS) and then proceed as for a liquid culture.
3. Once the smear has air dried, pass the smeared slide through a flame two or three times.
This kills the microbes in the smear and fixes the sample to the slide, be careful not to overheat the sample however as this can distort cellular morphology.
Gram stain procedure - Gram staining a sample
1. Gently flood the smear with crystal violet and leave for 1 minute. Tilt the slide slightly and gently rinse with tap water or distilled water.
Crystal violet is a water-soluble dye which enters the peptidoglycan layer in the bacterial cell wall.
2. Gently flood the smear with Gram’s iodine and leave for 1 minute. Tilt the slide slightly and gently rinse with tap water or distilled water. The smear will now appear purple.
Gram's iodine solution (iodine and potassium iodide) is added to form a complex with the crystal violet, which is much larger and is insoluble in water.
3. Decolorize the smear using 95% ethyl alcohol or acetone. Tilt the slide slightly and apply the alcohol drop by drop until the alcohol runs almost clear (5-10 seconds). Immediately rinse with water to avoid over-decolorizing.
Decolorizer dehydrates the peptidoglycan layer, shrinking and tightening it. In Gram positive bacteria, the large crystal violet-iodine complexes are then unable to penetrate and escape the thick peptidoglycan layer, resulting in purple stained cells. However, in Gram negative bacteria, the outer membrane is degraded, the thin peptidoglycan layer is unable to retain the crystal violet-iodine complexes and the color is lost.
4. Gently flood with safranin counterstain and leave for 45 seconds. Tilt the slide slightly and gently rinse with tap water or distilled water.
Safranin is weakly water soluble and will stain bacterial cells a light red, enabling visualization of Gram negative cells without interfering with the observation of the purple of the Gram positive cells.
5. Blot the slide dry on filter paper then view the smear using a light-microscope under oil-immersion.
Gram positive bacteria
Gram positive bacteria have a distinctive purple appearance when observed under a light microscope following Gram staining. This is due to retention of the purple crystal violet stain in the thick peptidoglycan layer of the cell wall. Examples of Gram positive bacteria include all staphylococci, all streptococci and some listeria species.
 Credit: Technology Networks
Credit: Technology Networks
Gram negative bacteria
Gram negative bacteria appear a pale reddish color when observed under a light microscope following Gram staining. This is because the structure of their cell wall is unable to retain the crystal violet stain so are colored only by the safranin counterstain. Examples of Gram negative bacteria include enterobacter species [June 6 2022], salmonella species and pseudomonas species.
Gram staining cannot be used reliably to assess bacterial phylogenetic relationships. The single membrane of Gram positive species is thought to be the ancestral state. Historically it had been thought that the second membrane found on Gram negative bacteria evolved just once and so all Gram negative species were more closely related to one another than they were to Gram positive species. However, genetic analysis has since shown this not to be the case and it is likely to have evolved multiple times in different lineages - a product of convergent evolution.
1. Gram, H.C. (1884). "Über die isolierte Färbung der Schizomyceten in Schnitt- und Trockenpräparaten". Fortschritte der Medizin (in German). 2: 185–189.
Correction: The article erroneously stated that enterococci were examples of Gram negative bacteria, this was updated on June 6, 2022 to correctly identify enterobacter species as examples of Gram negative bacteria.



