Viral Infections Predicted by Microscopy Deep Learning
Complete the form below to unlock access to ALL audio articles.
In humans, adenoviruses can infect the cells of the respiratory tract, while herpes viruses can infect those of the skin and nervous system. In most cases, this does not lead to the production of new virus particles, as the viruses are suppressed by the immune system. However, adenoviruses and herpes viruses can cause persistent infections that the immune system is unable to completely suppress and that produce viral particles for years. These same viruses can also cause sudden, violent infections where affected cells release large amounts of viruses, such that the infection spreads rapidly. This can lead to serious acute diseases of the lungs or nervous system.
Automatic detection of virus-infected cells
The research group of Urs Greber, Professor at the Department of Molecular Life Sciences at the University of Zurich (UZH), has now shown for the first time that a machine-learning algorithm can recognize the cells infected with herpes or adenoviruses based solely on the fluorescence of the cell nucleus. "Our method not only reliably identifies virus-infected cells, but also accurately detects virulent infections in advance," Greber says. The study authors believe that their development has many applications - including predicting how human cells react to other viruses or microorganisms. "The method opens up new ways to better understand infections and to discover new active agents against pathogens such as viruses or bacteria," Greber adds.
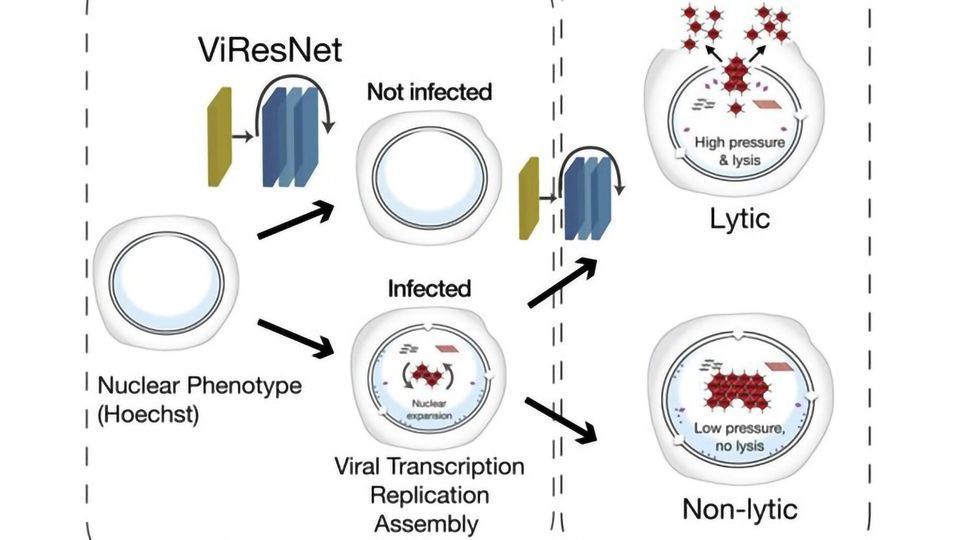
The analysis method is based on combining fluorescence microscopy in living cells with deep-learning processes. The herpes and adenoviruses formed inside an infected cell change the organization of the nucleus, and these changes can be observed under a microscope. The group developed a deep-learning algorithm - an artificial neural network - to automatically detect these changes. The network is trained with a large set of microscopy images through which it learns to identify patterns that are characteristic of infected or uninfected cells. "After training and validation are complete, the neural network automatically detects virus-infected cells," explains Greber.
Reliably predicting severe acute infections
The research team has also demonstrated that the algorithm is capable of identifying acute and severe infections with 95 percent accuracy and up to 24 hours in advance. Images of living cells from lytic infections, in which the virus particles multiply rapidly and the cells dissolve, as well as images of persistent infections, in which viruses are produced continuously but only in small quantities, served as training material. Despite the great precision of the method, it is not yet clear which features of infected cell nuclei are recognized by the artificial neural network to distinguish the two phases of infection. However, even without this knowledge, the researchers are now able to study the biology of infected cells in greater detail.
The group has already discovered some differences: The internal pressure of the nucleus is greater during virulent infections than during persistent phases. Furthermore, in a cell with lytic infection, viral proteins accumulate more rapidly in the nucleus. "We suspect that distinct cellular processes determine whether or not a cell disintegrates after it is infected. We can now investigate these and other questions," says Greber.
Reference: Andriasyan V, Yakimovich A, Petkidis A, et al. Microscopy deep learning predicts virus infections and reveals mechanics of lytic-infected cells. iScience. 2021;24(6). doi: 10.1016/j.isci.2021.102543
This article has been republished from the following materials. Note: material may have been edited for length and content. For further information, please contact the cited source.

