An Introduction to Post-Translational Modifications

Complete the form below to unlock access to ALL audio articles.
What are post-translational modifications?
Translation is often referred to as the "last step" of the "central dogma" of biology, whereby DNA is converted to RNA and then to protein. However, there are additional steps involved after protein synthesis that are necessary for a cell, tissue and organism to achieve its functional biology and diversity. Post-translational modifications are changes that are made to proteins after synthesis, typically mediated by enzymes.
Why are post-translational modifications important?
The sequencing of the first human genome in 2003 marked a major milestone in molecular biology. In the years since, it has become apparent that the human proteome – the collection of proteins that are or can be expressed in a human at a certain time – is categorically more complex.Why? The human proteome is extremely diverse. Whilst the genome is essentially constant across different cell populations in the human body (bar a few exceptions), in order for individual cells to perform their individual functions, and respond to environmental stimuli, a variety of different proteins must be expressed at different timepoints across cells, tissues and organs. It is estimated that there are ~20-25,000 protein-encoding genes, whilst the proteome is estimated to be over 1,000,000 proteins in size. The diversity of the proteome is achieved by several different mechanisms; post-translational modifications are one example.
 The process by which DNA is converted to proteins, and how post-translational modifications add to the complexity of the proteome. Credit: Technology Networks
The process by which DNA is converted to proteins, and how post-translational modifications add to the complexity of the proteome. Credit: Technology Networks
Post-translational modification examples
Here, we summarize five key examples of post-translational modifications. This is not an exhaustive list, as over 200 different types of post-translational modifications have been identified thus far.1Protein phosphorylation
Protein phosphorylation is the one of the most commonly occurring and most-studied post-translational modifications. It entails the phosphorylation of a specific amino acid residue through the addition of a phosphate group to a polar group R via a kinase, most commonly occurring at serine, tyrosine or threonine residues. The addition of the phosphate group results in a modification of the protein, whereby it transitions from being hydrophobic apolar to hydrophilic polar, enabling its interaction with other molecules – essentially "activating" it. A reversible post-translational modification, protein phosphorylation is important for cell regulation and the activation and deactivation of enzymes and receptors, which can be implicated in disease processes such as cancer.2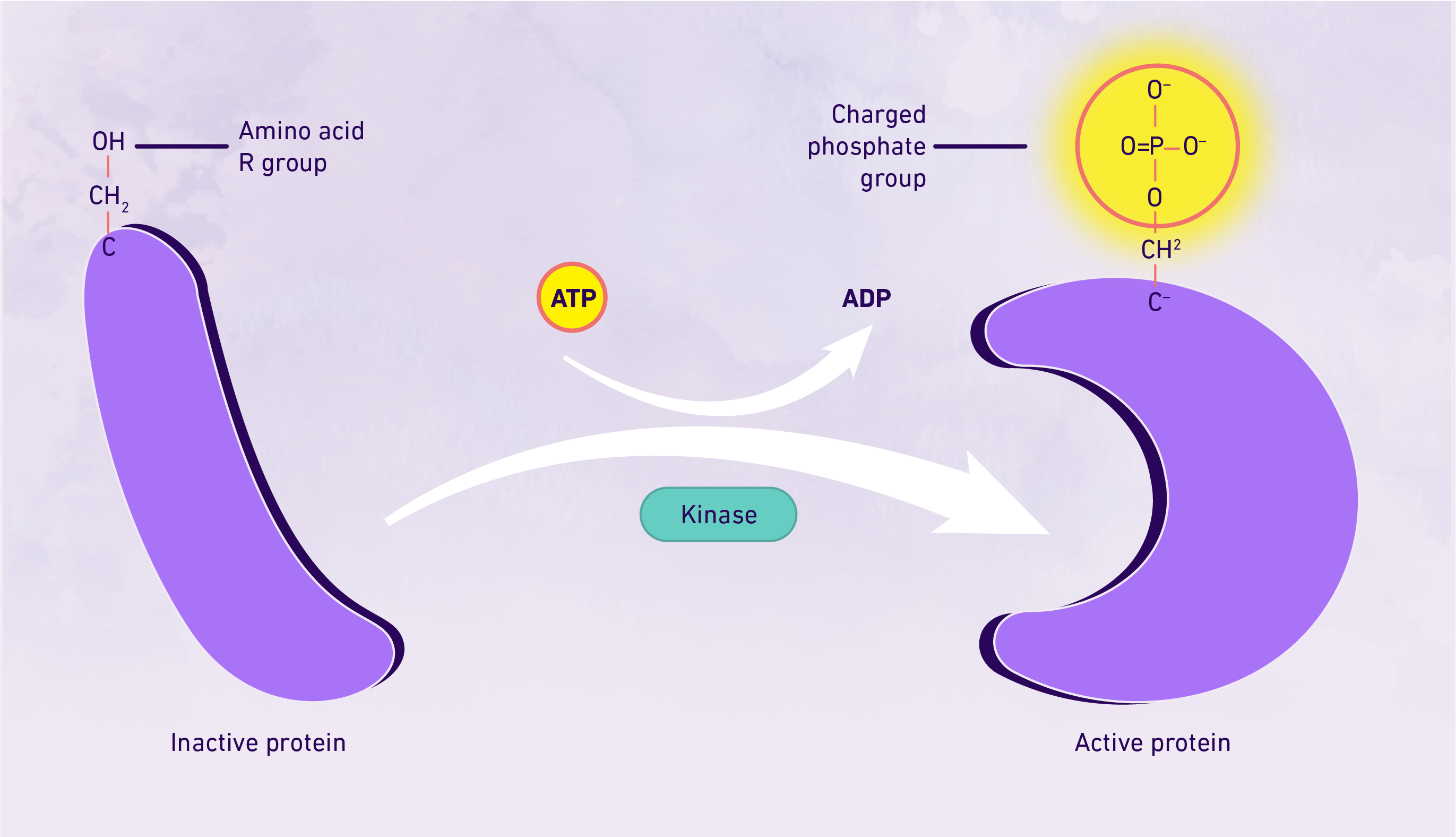 The key steps in protein phosphorylation summarized. Credit: Technology Networks
The key steps in protein phosphorylation summarized. Credit: Technology Networks
Protein glycosylation
Protein glycosylation is recognized as one of the most “complicated" yet most commonly occurring post-translational modifications.3 It involves the covalent addition of a carbohydrate moiety to an amino acid, forming a glycoprotein. Glycosylation reactions are diverse and catalyzed by various different enzymes, which attach specific glycans to specific amino acids. Glycoproteins are estimated to make up ~50% of the proteome; however, the study of the glycoproteome is challenging because of the large number and diversity of glycoprotein isoforms.4 Glycosylation of eukaryotic proteins is usually categorized into two major types; N-linked, whereby a sugar molecule is attached to the amide nitrogen of asparagine, and O-linked, where a sugar molecule is attached to the oxygen atom of serine or threonine.
There are an array of applications from glycoproteome research; many glycoproteins serve structural functions, whereas immunoglobulins are central to immunity and surface-presenting glycoproteins and glycolipids determine human blood group type.3 Thanks to advances in analytical technologies, such as mass spectrometry (MS), our knowledge of the human glycoproteome and the significance of glycosylation continues to grow.
Protein ubiquitination
Ubiquitin is a small protein – approximately 8kDa in size – that can bind to a substrate protein in a process known as ubiquitination, a type of post-translation modification that serves to regulate a protein's function or mark it for degradation. Ubiquitination occurs in three sequential steps that are catalyzed by three groups of enzymes. The key steps involved in protein ubiquitination. Credit: Technology Networks
The key steps involved in protein ubiquitination. Credit: Technology Networks
This process generally culminates with an isopeptide bond forming between ubiquitin and the lysine residue of the protein substrate.5 Monoubiquitination refers to the addition of one ubiquitin molecule, whereas the addition of several ubiquitin proteins is known as polyubiquitination.
Ubiquitination serves several functions, the most common being to flag proteins for degradation by the proteasome, but there are others including: immune and inflammatory response, organelle biogenesis and signaling roles in DNA repair.6
Protein methylation
Protein methylation has a regulatory role in many essential cellular processes, including gene transcription and signal transduction.7 In protein methylation, enzymes known as methyltransferases add a methyl group (most commonly donated by S-adenosyl-l-methionine, or SAM) to specific amino acids on a protein molecule, such as the lysine and arginine residues. Protein methylation can have effects on:- Protein stability
- Protein subcellular localization
- Protomer binding affinity
- Protein-protein interactions
- Other protein modification events
As such, it is often studied in the context of cancer where the regulation of cellular processes and cell cycle regulation can go awry.8 The characterization of individual protein methylation processes is a growing area of research and is gradually becoming easier to study thanks to advances in MS; however, the complexity of the methylome makes it challenging to profile as a collective using the currently available tools.9
Protein acetylation
Protein acetylation is a common post-translational modification in eukaryotes and involves the addition of an acetyl group to nitrogen via reversible and irreversible processes.
Acetylation has been studied largely in histones, the proteins that pack DNA into chromosomes, where the acetylation of the lysine side chain (ε-NH2) on the N-termus of histones is associated with the regulation of gene expression. If lysine is acetylated, it is no longer positively charged. In turn, the binding of DNA to the histone is relaxed, which facilitates the transcription of genes. Whilst histone research has dominated the field, scientists now suspect that there are a wide range of protein acetylations which could carry vital physiological functions across different biological processes, particularly in bacteria.10,11
 An image detailing the acetylation of a lysine side chain. Credit: Technology Networks
An image detailing the acetylation of a lysine side chain. Credit: Technology Networks
Post-translational modification research outlooks
Post-translational modifications are key to the diversity of the proteome, and in turn, the diversity of an organism. Whilst MS-based proteomics has facilitated novel insights on the role post-translational modifications play in biological processes, we still have a way to go yet. In the future, characterizing and mapping the modified proteome with high sensitivity and specificity will undoubtedly unlock major advances in scientific disciplines such as diagnostics and drug development in our pursuit towards personalized medicine.12Post-translational modification summary table
Post-translation modification | Mechanism |
Protein phosphorylation | Addition of a phosphate group to an amino acid residue. |
Protein glycosylation | Covalent addition of a carbohydrate moiety to an amino acid, forming a glycoprotein. |
Protein ubiquitination | Binding of a ubiquitin protein to a protein via a three-step process. |
Protein methylation | Addition of a methyl group, most often at lysine or arginine residues. |
Protein acetylation | Addition of an acetyl group to an N-terminus of a protein, or at lysine residues. |
1. Minguez P, Parca L, Diella F, et al.
Deciphering a global network of functionally associated post-translational
modifications. Mol Syst Biol. 2012;8:599. doi: 10.1038/msb.2012.31.
2. Ardito F, Giuliani M, Perrone D, Troiano G, Lo Muzio L. The crucial role of protein phosphorylation in cell signaling and its use as targeted therapy (Review). Int J Mol Med. 2017;40(2):271-280. doi: 10.3892/ijmm.2017.3036.
3. Spiro RG. Protein glycosylation: nature, distribution, enzymatic formation, and disease implications of glycopeptide bonds. Glycobiology. 2002;12(4):43R-56R. doi: 10.1093/glycob/12.4.43R.
4. Gill DJ, Clausen H, Bard F. Location, location, location: new insights into O-GalNAc protein glycosylation. Trends Cell Biol. 2011;21(3):149-158. doi: 10.1016/j.tcb.2010.11.004.
5. Komander D, Rape M. The ubiquitin code. Annu Rev Biochem. 2012;81(1):203-229. doi: 10.1146/annurev-biochem-060310-170328.
6. Sun L, Chen ZJ. The novel functions of ubiquitination in signaling. Curr Opin Cell Biol. 2004;16(2):119-126. doi: 10.1016/j.ceb.2004.02.005.
7. Murn J, Shi Y. The winding path of protein methylation research: milestones and new frontiers. Nat Rev Mol Cell Biol. 2017;18(8):517-527. doi: 10.1038/nrm.2017.35.
8. Hamamoto R, Nakamura Y. Dysregulation of protein methyltransferases in human cancer: An emerging target class for anticancer therapy. Cancer Sci. 2016;107(4):377-384. doi: 10.1111/cas.12884.
9. Luo M. Chapter 10 - Current methods for methylome profiling. In: Zheng YG, ed. Epigenetic technological applications. Academic Press; 2015:187-217. doi: 10.1016/B978-0-12-801080-8.00010-7.
10. Xia C, Tao Y, Li M, Che T, Qu J. Protein acetylation and deacetylation: An important regulatory modification in gene transcription (Review). Exp Ther Med. 2020;20(4):2923-2940. doi: 10.3892/etm.2020.9073.
11. Christensen DG, Xie X, Basisty N, et al. Post-translational protein acetylation: an elegant mechanism for bacteria to dynamically regulate metabolic functions. Front Microbiol. 2019;10:1604. doi: 10.3389/fmicb.2019.01604.



