Electron Relationships in Alzheimer’s Protein Found to Be “Outside the Common Sense of Organic Chemistry”
Complete the form below to unlock access to ALL audio articles.
The chemical interactions that give proteins their shape may be weaker and more numerous than previously recognized. These weak connections provide a new way for researchers to understand proteins that cause disease and help them gain insights into the fundamentals of chemistry.
Chemists at the University of Tokyo modeled the building blocks of the protein structure that causes Alzheimer's disease, amyloid beta sheets. Their calculations revealed that some atoms too far apart to bond were still in each other's "electron neighborhoods."
"This is so strange. It's outside the common sense of organic chemistry," said Professor Tomohiko Ohwada from the University of Tokyo.
Atoms - the building blocks of life represented in the periodic table of the elements - connect together to build molecules by sharing or stealing electrons from other atoms. Those electrons fly around super-fast in a cloud of electrons.
The traditional understanding of chemical bonds is that in large molecules, atoms must be side by side to share electrons.
Ohwada's research team calculated that atoms located far apart do not share electrons, but their electron clouds can still influence each other. They call this influence "through-space bond path interactions." The through-space bond path interactions are extremely weak, but common enough to add up to a potentially significant influence on the overall structure of large molecules.
"We studied the amyloid beta structure because everyone knows it can cause disease, but nobody really knows how the problematic structure develops," said Ohwada.
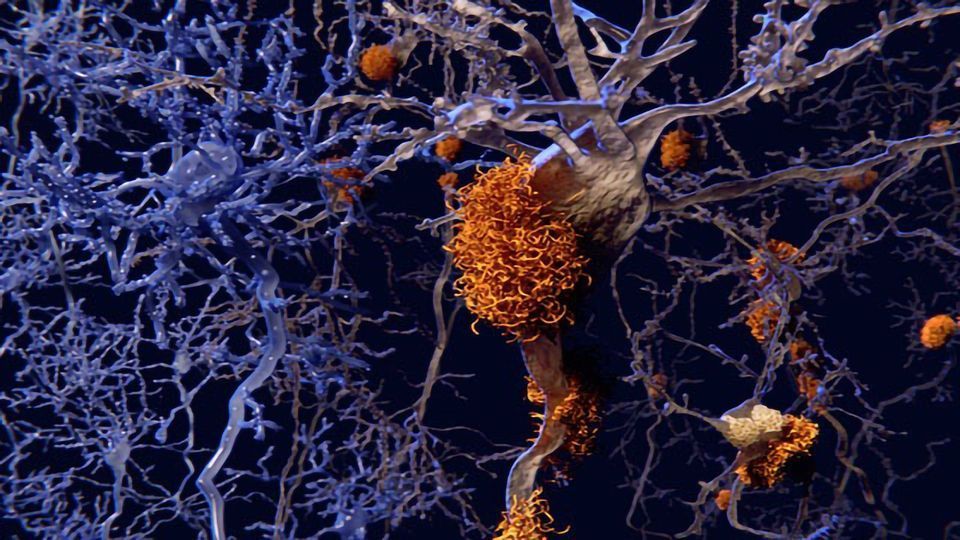
Abnormal accumulation of amyloid beta sheets may cause Alzheimer's disease or cancers.
The beta sheet structure refers to long chains of protein folded at regular intervals and stacked one on top of another into a flat sheet. Through-space bond path interactions within and between the protein chains likely stabilize the structure and may help it clump together into disease-causing plaques.
Recognizing the location and nature of through-space bond path interactions may help researchers predict the true structure and behavior of a molecule based only on its chemical sequence.
The research team has so far only studied through-space bond path interactions in the water-repelling portions of synthetically built mini amyloid beta sheets. They plan to expand their computations to the water-attracting portions and to larger molecules.
"In theory, it might be possible to build an artificial molecule that could form through-space bond path interactions with natural proteins and change their activity," said Ohwada.
The computations in this research were performed at the Research Center for Computational Science in Okazaki, Japan.
Reference: Luhan Zhai, Yuko Otani and Tomohiko Ohwada. 2019. Uncovering the Networks of Topological Neighborhoods in β-Strand and Amyloid β-Sheet Structures. Scientific Reports. DOI: https://doi.org/10.1038/s41598-019-47151-2
This article has been republished from the following materials. Note: material may have been edited for length and content. For further information, please contact the cited source.

