Transcriptional Regulation of Memory Formation in the Hippocampus

Complete the form below to unlock access to ALL audio articles.
MIT neuroscientists have uncovered a cellular pathway that allows specific synapses to become stronger during memory formation. The findings provide the first glimpse of the molecular mechanism by which long-term memories are encoded in a region of the hippocampus called CA3.
The researchers found that a protein called Npas4, previously identified as a master controller of gene expression triggered by neuronal activity, controls the strength of connections between neurons in the CA3 and those in another part of the hippocampus called the dentate gyrus. Without Npas4, long-term memories cannot form.
“Our study identifies an experience-dependent synaptic mechanism for memory encoding in CA3, and provides the first evidence for a molecular pathway that selectively controls it,” says Yingxi Lin, an associate professor of brain and cognitive sciences and a member of MIT’s McGovern Institute for Brain Research.
Lin is the senior author of the study, which appears in the Feb. 8 issue of Neuron. The paper’s lead author is McGovern Institute research scientist Feng-Ju (Eddie) Weng.
Synaptic strength
Neuroscientists have long known that the brain encodes memories by altering the strength of synapses, or connections between neurons. This requires interactions of many proteins found in both presynaptic neurons, which send information about an event, and postsynaptic neurons, which receive the information.
Neurons in the CA3 region play a critical role in the formation of contextual memories, which are memories that link an event with the location where it took place, or with other contextual information such as timing or emotions. These neurons receive synaptic inputs from three different pathways, and scientists have hypothesized that one of these inputs, from the dentate gyrus, is critical for encoding new contextual memories. However, the mechanism of how this information is encoded was not known.
In a study published in 2011, Lin and colleagues found that Npas4, a gene that is turned on immediately following new experiences, appears to act as a master controller of the program of gene expression required for long-term memory formation. They also found that Npas4 is most active in the CA3 region of the hippocampus during learning. This activity was already known to be required for fast contextual learning, such is required during a type of task known as contextual fear conditioning. During the conditioning, mice receive a mild electric shock when they enter and explore a specific chamber. Within minutes, the mice learn to fear the chamber, and the next time they enter it, they freeze.
When the researchers knocked out the Npas4 gene, they found that mice could not remember the fearful event. They also found the same effect when they knocked out the gene just in the CA3 region of the hippocampus. Knocking it out in other parts of the hippocampus, however, had no effect on memory.
In the new study, the researchers explored in further detail how Npas4 exerts its effects. Lin’s lab had previously developed a method that makes it possible to fluorescently label CA3 neurons that are activated during this fear conditioning. Using the same fear conditioning process, the researchers showed that during learning, certain synaptic inputs to CA3 neurons are strengthened, but not others. Furthermore, this strengthening requires Npas4.
The inputs that are selectively strengthened come from another part of the hippocampus called the dentate gyrus. These signals convey information about the location where the fearful experience took place.
Without Npas4, synapses coming from the dentate gyrus to CA3 failed to strengthen, and the mice could not form memories of the event. Further experiments revealed that this strengthening is required specifically for memory encoding, not for retrieving memories already formed. The researchers also found that Npas4 loss did not affect synaptic inputs that CA3 neurons receive from other sources.
Kimberly Raab-Graham, an associate professor of physiology and pharmacology at Wake Forest University School of Medicine, says the researchers used an impressive variety of techniques to unequivocally show that contextual memory formation is tightly controlled by Npas4.
“The major finding of the study is that contextual memory is driven by a single circuit and comes down to a single transcription factor,” says Raab-Graham, who was not involved in the study. “When they knocked out the transcription factor, they removed contextual memory formation, and they could restore it by adding the transcription factor.”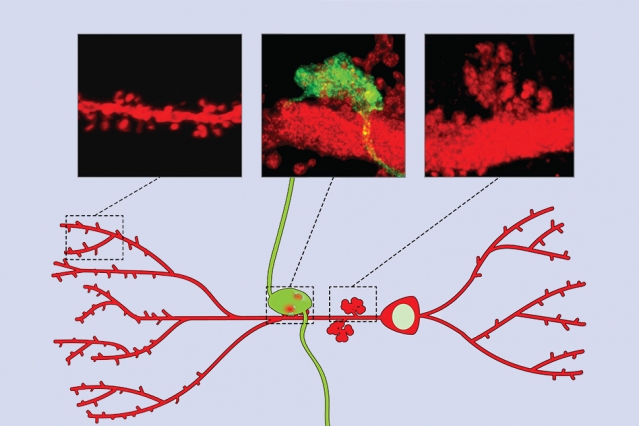 This schematic shows two types of synaptic structures on CA3 neurons. The smaller spines, labeled in red in the image at left, are more involved in memory retrieval. The large green structure (center image) represents a synapse between a CA3 neuron and an incoming axon from the dentate gyrus. At right is a close-up of a large, multi-branched spine without the labeled input from the dentate gyrus. Credit: Lin Lab, MIT.
This schematic shows two types of synaptic structures on CA3 neurons. The smaller spines, labeled in red in the image at left, are more involved in memory retrieval. The large green structure (center image) represents a synapse between a CA3 neuron and an incoming axon from the dentate gyrus. At right is a close-up of a large, multi-branched spine without the labeled input from the dentate gyrus. Credit: Lin Lab, MIT.
Synapse maintenance
The researchers also identified one of the genes that Npas4 controls to exert this effect on synapse strength. This gene, known as plk2, is involved in shrinking postsynaptic structures. Npas4 turns on plk2, thereby reducing synapse size and strength. This suggests that Npas4 itself does not strengthen synapses, but maintains synapses in a state that allows them to be strengthened when necessary. Without Npas4, synapses become too strong and therefore cannot be induced to encode memories by further strengthening them.
“When you take out Npas4, the synaptic strength is almost saturated,” Lin says. “And then when learning takes place, although the memory-encoding cells can be fluorescently labeled, you no longer see the strengthening of those connections.”
In future work, Lin hopes to study how the circuit connecting the dentate gyrus to CA3 interacts with other pathways required for memory retrieval. “Somehow there’s some crosstalk between different pathways so that once the information is stored, it can be retrieved by the other inputs,” she says.
This article has been republished from materials provided by MIT. Note: material may have been edited for length and content. For further information, please contact the cited source.
Reference:
Feng-Ju Weng, Rodrigo I. Garcia, Stefano Lutzu, Karina Alviña, Yuxiang Zhang, Margaret Dushko, Taeyun Ku, Khaled Zemoura, David Rich, Dario Garcia-Dominguez, Matthew Hung, Tushar D. Yelhekar, Andreas Toft Sørensen, Weifeng Xu, Kwanghun Chung, Pablo E. Castillo, Yingxi Lin. Npas4 Is a Critical Regulator of Learning-Induced Plasticity at Mossy Fiber-CA3 Synapses during Contextual Memory Formation. Neuron, 2018; DOI: 10.1016/j.neuron.2018.01.026

