Antigenic Drift vs Antigenic Shift
Even with careful research, a certain amount of educated guesswork is involved for manufacturers who select strains to incorporate into the annual flu vaccine.

Complete the form below to unlock access to ALL audio articles.
As part of the host-pathogen arms race, viruses are continually evolving to evade the host immune response, be it from previous infection or immunity acquired through vaccination.
Many people get their annual flu shots in the hope it will protect them from that year’s seasonal flu outbreaks. However, there is no guarantee that the strains predicted to be circulating that year will actually be the ones that do. Even with careful research and probability modeling, a certain amount of educated guesswork is involved for manufacturers who select strains to incorporate into the annual vaccine. And the main culprit for their headaches? Antigenic drift and antigenic shift.
Contents
Antigenic drift vs antigenic shift: table
What are the consequences of antigenic drift vs antigenic shift?
What is antigenic drift?
Antigens are molecules that are recognized by the host immune system as foreign and induce an immune response. Consequently, they will often be surface proteins, like haemagglutinin (HA) or neuraminidase (NA) in the case of the influenza virus.
Antigenic drift is a natural process whereby mutations (mistakes) occur during replication in the genes encoding antigens that produce alterations in the way they appear to the immune system (antigenic changes) (Figure 1). Not all genetic mutations will result in antigenic changes depending on 1) their position in the triplet code (non-coding changes) or 2) if the change they produce does not affect the region of the protein recognized by the immune system.
If the changes reduce virus survival, they will be selected against and lost from the population. If they have no effect or offer benefits, such as immune evasion, they are likely to persist and may even spread through the population, depending on other associated changes.
Even if a host has not encountered that particular virus before, partial immunity may be gained from infection or vaccination with a closely related virus in the past.
Figure 1: Representation of antigenic drift producing alterations in how the virus may appear to the immune system. Credit: Technology Networks.
What is antigenic shift?
 Figure 2: Structure of the influenza A virus. Credit: Technology Networks.
Figure 2: Structure of the influenza A virus. Credit: Technology Networks.
Influenza has a negative sense single stranded RNA genome, encapsulated by nuclear protein, which consists of eight segments (Figure 2). Due to its segmented nature, influenza viruses can swap whole sections of their genome. If the segment swapped encodes an influenza antigen (such as HA or NA) which is targeted by the host immune system, this is termed antigenic shift and can radically alter a host immune system’s ability to recognize the virus (Figure 3).
Influenza A is responsible for most seasonal and pandemic flu outbreaks in humans. As the primary surface antigens, HA and NA types are incorporated into the naming scheme for Influenza A virus strains (e.g., A/Perth/16/2009 (H3N2)). When antigenic shift occurs, it is possible that an H1N1 virus and H3N2 virus may interact to produce an H1N2 virus and an H3 N1 virus, for example.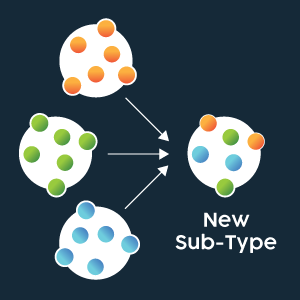
Figure 3: Representation of genome segment re-assortment (antigenic shift) between different strains producing a novel virus. Credit: Technology Networks.
Antigenic drift vs antigenic shift: table
Table 1: Features of antigenic drift and antigenic shift.
| Feature | Antigenic drift | Antigenic shift |
| Mechanism | Small mutations during replication lead to changes in the genes encoding antigens. | Viruses swap whole sections of their genomes, leading to changes in antigen genes. |
| Changes to genome | Small or point mutations. | Large changes to sections of genome. |
| Result | Can help viruses gradually evolve and evade immune systems. | Can produce entirely new virus strains. |
What are the consequences of antigenic drift vs antigenic shift?
Individuals may have partial immunity, from previous infection or vaccination, to infection with influenza strains that have undergone antigenic drift. However, antigenic shift generates novel lineages to which immunity is therefore often very poor across the population. Consequently, pandemics are nearly always a result of antigenic shift events.
Pandemic outbreaks have been recorded through history, the more recent of which have been attributed to specific subtypes (summarized in Table 1).
Influenza virus surface protein types determine the types of receptors that the virus can interact with, and consequently, the host species which can be infected.
Influenza viruses are known to transmit zoonotically as some strains can infect multiple species. Pigs are a common intermediate host for influenza strains between humans, who have α2,6 receptors in their respiratory tract, and species such as birds who have α2,3 receptors. This is because pigs are the only mammalian species to have both receptor types and are therefore susceptible to and enable productive replication of avian and human influenza viruses in one animal. This also means pigs are a common mixing pot for cases of antigenic shift.
A highly pathogenic strain that can infect birds but is unable to infect humans may infect an intermediate host, like a pig, at the same time as a low-pathogenicity strain that can infect humans. The highly pathogenic strain may swap segments with the low-pathogenicity strain and voila – a highly pathogenic strain that can infect people, cue a new pandemic!
Table 2: Influenza pandemics in humans.
| Date | Name of pandemic/description | Estimated deaths | Subtype involved |
| 1510 | Starts in Africa, proceedes northward to all of Europe and the Baltic States. Attack rates are extremely high, but fatality is low and said to be restricted to young children. | Low | ? |
| 1557-1558 | The first influenza pandemic in which global involvement and westward spread from Asia to Europe documented. Highly fatal, with deaths recorded as being due to "pleurisy and fatal peripneumony". High mortality in pregnant women also recorded. | High | ? |
| 1580 | Influenza pandemic originates in Asia, spreading to Africa, and then Europe. Illness rates high. | 8000 in Rome, Spanish cities decimated | ? |
| 1729-1730 | Worldwide Influenza Pandemic. | ? | ? |
| 1761-1762 | Originates in the Americas, spreads to Europe and around the globe in 1762. First pandemic to be studied by multiple observers who communicate with each other in learned societies and through medical journals and books. Influenza is characterized clinically to a greater degree than it has been previously, as physicians carefully record observations on series of patients and attempt to understand what would later be called the pathophysiology of the disease. | ? | ? |
| 1780-1782 | Worldwide Flu. | ? | ? |
| 1830-1833 | Started in China, spread southwards by sea to the Philippines, India and Indonesia, and across Russia into Europe. Reaches the Americas by 1831. Overall the attack rate is estimated at 20–25% of the population, but the mortality rate is not exceptionally high. | Not exceptionally high | ? |
| 1857-1859 | Worldwide Flu Pandemic. | ? | ? |
| 1889–1890 | Asiatic or Russian Flu. | 1 million | H3N8 or H2N2 |
| 1918-1920 | Spanish Flu. | 20 to 100 million | H1N1 |
| 1957-1958 | Asian Flu. | 1 to 1.5 million | H2N2 |
| 1968-1969 | Hong Kong Flu - over 65's most vulnerable. | 0.75 to 1 million | H3N2 |
| 1977–1978 | Russian Flu. | - | H1N1 |
| 2009-2010 | Swine Flu - Originated in Mexico. The virus, a combination of a Eurasian swine flu virus with another strain that was itself a mix of bird, swine and human flu virus. | 18,000 to 284,500 | H1N1/09 |





