High Content Image-based siRNA Screen Identifies Kinases Important for Haemostasis

Complete the form below to unlock access to ALL audio articles.
The endothelial cells that line our blood vessels play an important role in maintaining and regulating blood flow, contributing to the natural process called haemostasis. They are also involved in the clotting cascade initiated when a blood vessel is damaged. One mechanism by which endothelial cells carry out their role is by secreting factors into the bloodstream that help recruit the blood clotting factors and wound-healing cells to a wound.
Surprisingly, defects in blood clotting are among the most common disorders in humans. In fact, at least 1% of the World’s population is thought to suffer from the most common of these disorders, von Willebrand disease. The disease is caused by a mutation in the gene that codes for the von Willebrand factor, a blood protein that acts like a glue to stick platelets together and heal wounds.
Read more about von Willebrand factor research: https://www.technologynetworks.com/tn/news/engineered-clotting-protein-stops-bleeding-in-most-common-inherited-bleeding-disorder-193423
High Content Imaging
Research by Janos Kriston-Vizi’s group at the MRC’s Laboratory for Molecular Cell Biology at UCL, London, has used a novel approach to understand which genes regulate the function and release of such factors from endothelial cells. As Janos explains, “We used a novel high-content screening method to visualize endothelial cells and one of the key factors they release that contributes to hemostasis, von Willebrand factor, by microscopy.”
The team achieved this by developing a workflow for high throughput measurement of the size and length of Weibel-Palade Bodies, secretory vesicles that contain von Willebrand factor, in primary human umbilical vein endothelial cells (HUVEC).

Image analysis workflow used to label and measure Weibel-Palade Bodies size and length.
Their unique workflow enabled them to disrupt over 800 genes and analyse the phenotypic effect of these disruptions on von Willibrand factor release from endothelial cells.
The 800 candidate genes mainly encoded protein kinases that were thought to regulate von Willebrand factor formation. The team used electroporation to deliver siRNA to umbilical vein endothelial cells to knockdown kinase expression, and observed the effect on von Willebrand factor release. The experiments generated 10,560 images and were performed using a Perkin Elmer Opera system. The image data was analysed using freely available software. Their experimental approach measured more than 10million Weibel-Palade Bodies.
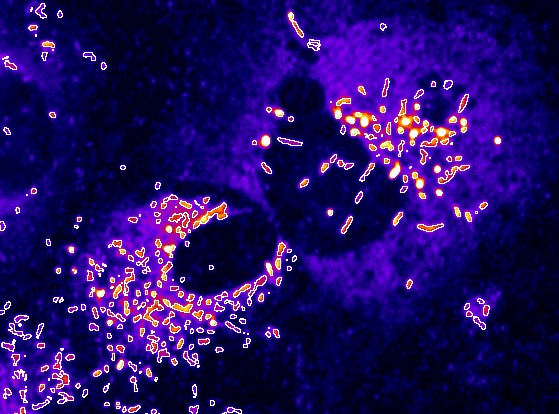
Automated measurement of Weibel-Palade Bodies, which contain von Willebrand factor.
Open Source
“These images contain a large amount of data that will not only be useful for researchers specifically interested in von Willebrand disease, but also more generally for the wider research community.” Explained Janos, before adding, “Sharing that amount of large image data is challenging, however, due to the traditional printed format of scientific journals.”
To overcome the problem of sharing this large dataset in a meaningful way, the group made their large image set available at the Harvard Dataverse repository. They also describe their findings and open source image analysis workflow in their latest paper, published in the Functional Genomics collection of the journal, Scientific Data (Ketteler et al. 2017).
Reference:
Ketteler, R., Freeman, J., Ferraro, F., Bata, N., Cutler, D. and Kriston-Vizi, J. (2017). Image-based siRNA screen to identify kinases regulating Weibel-Palade body size control using electroporation. Scientific Data, 4, p.170022.



