Metabolomics-Based Blood Test Could Enable Earlier Detection of Cancer
Complete the form below to unlock access to ALL audio articles.
Along with the development of new therapies, advances in technologies that enable earlier diagnosis of cancer are critical to improve patient outcomes. Several screening programs and referral pathways are enabling timely identification of cancer in individuals presenting with symptoms indicative of a specific cancer. But for patients with more general symptoms, such as fatigue and unexplained weight loss, the journey to diagnosis can be much longer. Better methods are urgently needed to identify cancer in these patients sooner and reduce the negative impacts associated with late-stage cancer diagnoses.
In a study recently published in Clinical Cancer Research, a team of scientists demonstrate the potential of nuclear magnetic resonance (NMR)-based metabolomics to detect cancer in patients with nonspecific symptoms, as well as determine whether a cancer is metastatic. The approach was shown to be highly sensitive and specific, identifying patients with solid tumors with 94% sensitivity and 82% specificity. In the study, the blood of 294 patients was analyzed, and differences between the metabolic “fingerprints” of those with and without cancer, and with and without metastases were detected. Importantly, in two patients, metabolomics analysis of blood was able to detect a new or recurring cancer earlier than conventional imaging methods.
Technology Networks had the pleasure of speaking to first author, Dr. James Larkin, to learn more about the study, the benefits of biofluid metabolomics and how an NMR-based approach could help to improve the way that cancer is detected and monitored in patients with nonspecific symptoms.
Anna MacDonald (AM): Can you tell us about some of the limitations of current cancer referral pathways? What options are available for patients presenting with nonspecific symptoms?
James Larkin (JL): Most current cancer referral pathways are designed for people presenting with specific symptoms. Good examples would be if you have a persistent cough as a symptom of potential lung cancer, or if you have a noticeable lump that’s recently grown. Patients with nonspecific symptoms, like weight loss or fatigue, are difficult to diagnose because there’s many potential causes and it’s difficult for GPs to accurately refer patients for further investigations. In a lot of cases, patients are asked to monitor their symptoms and return if they get worse which can mean that if they do have a cancer, it’s growing more.
This has been recognized as a problem and there’s been a shift recently in the UK, with a drive to introduce nonspecific symptom pathways. These are being rolled out nationwide over the coming years, but they still have the problem that many patients with nonspecific symptoms don’t actually have cancer. What’s needed is a test like ours that can accurately and rapidly tell if a patient with nonspecific symptoms has a cancer. This would allow GPs to quickly refer patients for imaging, with the confidence that they’re not sending people unnecessarily.
AM: What led you to investigate biofluid metabolomics as an alternative approach for this group of patients?
JL: We have always been interested in cancer diagnosis, but this specific approach was inspired by our previous work in other diseases. We were studying inflammatory disease like multiple sclerosis and trying to work out how the body responds to these diseases. As we worked, it became clear that one of the effects was a change in the immune system response. We suspected that this immune system change would have effects on the way that the rest of the body metabolizes. We began exploring this change in metabolism and discovered that you can use patterns of change to predict when multiple sclerosis patients transition from the earlier “relapsing-remitting” to the later “secondary progressive” form of the disease.
It’s well-known that cancer involves a strong immune system response and also involves changes in metabolism. This led to us hypothesizing that we might be able to identify and monitor cancers using the same approach. We began looking at just secondary brain tumors, but soon expanded to looking at cancer more generally because of the pressing clinical need.
AM: Can you briefly talk me through the NMR metabolomics analysis?
JL: Sure. There’s three parts to the process. The first is blood collection, the second is the NMR analysis itself and the third is the computational analysis of the NMR data.
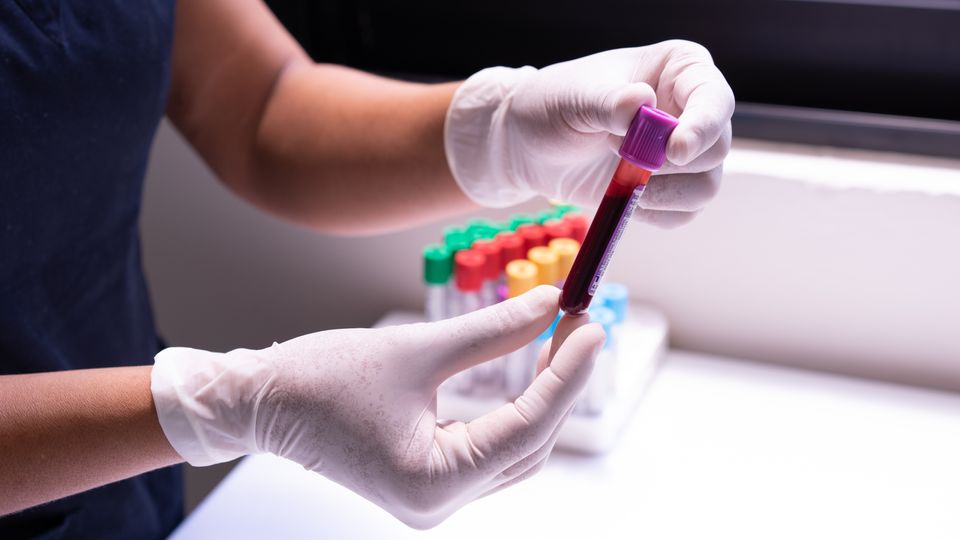
The first step, the blood sample, is routine and is just like any other blood test – a needle is put in a vein, often in your arm, and we collect a blood sample in a regular blood collection tube. We need around 5mL of blood, about a teaspoon-full. The blood is then spun in a centrifuge to separate the red blood cells from the clear portion of the blood, the serum. We use the serum for the NMR analysis.
The NMR analysis uses a machine called a nuclear magnetic resonance spectrometer. It’s a large machine that uses high magnetic fields and radio waves to look at the composition of a sample. It works because all molecules have a particular frequency that they resonate at, much like a tuning fork resonates at one specific frequency. The NMR analysis works much like striking thousands of tuning forks at once then carefully listening to the sound to work out which tuning forks are present. This gives us a data-rich “spectrum” which contains information on all the small molecules, or metabolites, present in the blood serum.

Credit: James Larkin
The third and final step is the computational analysis. We use machine learning and other statistical techniques to look at the pattern of metabolites present in a patient’s blood sample. We discovered that there are some patterns that are more likely in cancer and some patterns that are more likely if you have a non-cancer disease. We can use these changing patterns to determine whether a patient has a cancer or not.
AM: In your study, you demonstrated that metabolomic biomarkers could be used to identify cancer in patients with nonspecific symptoms. Can you tell us about some of the main metabolic changes observed between the groups in the study? What could be responsible for these changes?
JL: This is an interesting point. The reason we carried out the work was to see if we could use our technology to diagnose cancer. For diagnosis to work, all that matters is “does the pattern of metabolites in the patient’s blood closely match the pattern found in cancer patients or non-cancer patients’ blood?”. This means that from a diagnostic point of view, we don’t need to understand, or even look at, the changes in individual metabolites that we see.
Of course, the changes in specific metabolites do have immense importance in trying to understand the underlying cancer. We’ve speculated a little bit on why these changes have happened in the paper, but broadly we’ve seen decreases in fatty acids and increases in glucose and some amino acids in cancer patients. We suspect that what we’re seeing is the body’s response to the presence of the cancer, not just the cancer itself. Metabolism is normally very tightly controlled and so changes that can be detected are almost certain to involve parts of the body like the liver.
AM: You were also able to distinguish between metastatic and non-metastatic cancers. How is this information usually obtained? What difference could a blood test that enables metastatic status to be determined make to patient pathways?
JL: Normally the presence of metastases is confirmed either by imaging, such as positron emission tomography (PET), or by biopsy, that is removing something like the local lymph nodes and examining them under a microscope. Deciding which patients need investigation is a matter for the oncologist and will depend on an individual’s risk of developing metastases. This process isn’t perfect and right now many patients without metastases are sent for imaging and many patients with metastases aren’t sent for imaging soon enough. A blood test that identifies metastases would allow a much better identification of those patients that need to go for imaging, allowing resources to be prioritized and more patients with metastases identified sooner. Being able to identify metastases is important because there are some therapies, like immunotherapies, that are most beneficial in patients with metastases. Like all cancer treatments, earlier is better.
AM: In two cases, cancer was identified by metabolomic analysis before it was detected by conventional imaging. How could metabolomics be applied in the clinic to support earlier cancer diagnosis?
JL: There’s a number of ways we can see metabolomics helping. Firstly, patients with nonspecific symptoms can benefit as I said earlier. Beyond that, there’s lots of examples where we can see potential benefit. Consider relapse monitoring – after a patient finishes a course of treatment for a cancer they will be followed up for some years to make sure that the cancer doesn’t return.
A second example would be patients who have non-cancer conditions that may develop into cancers, for example a benign lump is likely to stay benign, but may develop into a cancer. Many of these conditions have monitoring pathways in place, but a lot of patients that are passing through them will never go on to develop a cancer. This makes the pathways inefficient and potentially costly. Metabolomic analysis may provide an “early warning” of sorts that the benign condition may be turning cancerous.
AM: How does this approach compare to other liquid biopsy approaches such as ctDNA analysis? What benefits does it offer?
JL: Broadly speaking, other approaches to cancer detection work by either looking for something specific released by the tumor, like a specific blood marker or your example of circulating tumour DNA, or by physically looking for a tumor, perhaps using an imaging approach like MRI or CT.
Our approach is different because we are looking at a range of small metabolites, none of which are specific to the tumor alone. All people have these metabolites circulating all the time and they’re essential to life. What’s unique about our approach is that we’re looking only at the pattern of these metabolites and seeing how the pattern itself is changing in cancer. We strongly suspect that our signal is being “amplified” by the rest of the body which is giving us an advantage in terms of our detection power. Also, by using a regular blood sample, we’re hoping that we will be able to deploy our test in the widest possible number of locations.
AM: Lead author of the study, Dr. Fay Probert, commented in a press release that “The goal is to produce a test for cancer that any GP can request.” What steps are needed before this can be achieved? Would the test remain broad, detecting a range of cancers, or do you envisage several tests being developed for the detection of specific cancers?
JL: Like all medical diagnostic tests, we must take our technology from its current form and put it together as a marketable product with regulatory approval. This is a complex process and we are setting up a spinout company from the University of Oxford to carry out this work. At the simplest level, we will need to conduct a larger trial with more patients that demonstrates that our technology is reliable in a broad population, is safe, and is effective at finding cancer. The body of evidence we gather will be sent to the medical regulators for approval.
Downstream, we anticipate a number of tests, each of which is specific to a particular cancer. However, for now the strongest evidence we have is for our broad test that detects a range of cancers. The clever part of our analysis is that as we recruit more patients to trials, we will be able to develop better diagnostic tests using data from those patients. This should greatly increase the speed at which we can develop new specific tests.
Dr. James Larkin was speaking to Anna MacDonald, Science Writer for Technology Networks.




