Discrimination of Meat Species Using Raman Spectroscopy and PCA
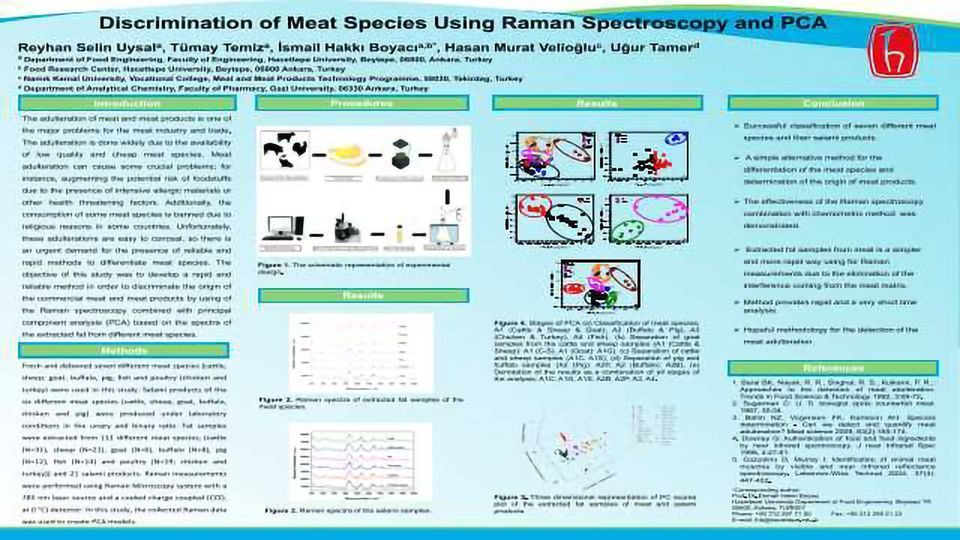
Differentiation the origin of the meat species and meat products was successfully performed based on their extracted fat samples using Raman spectroscopy and the principal component analysis (PCA). Extracted fat samples which were obtained from different meat species (cattle, sheep, goat, buffalo, pig, fish, poultry (chicken and turkey)) and their salami products were assayed based on their Raman measurements of their fat profiles. The collected Raman data were analyzed with a four-stage PCA . Different combinations of pre-processing techniques (baseline correction, different orders of derivatives, mean centre, normalising, smoothing, and auto-scaling) were applied to Raman data before the creating of the models to enhance the performance and the decomposition ability of the model. Four different PCA models were created with a total of 132 Raman data of the samples. The principal component (PC) scores were used to plot the score graphs of the samples and to create the clusters of same meat species in the same area. Seven meat species and their salami products were successfully differentiated from each other according to their origin. The results showed that the method that is Raman spectroscopy with PCA is practicable and usable analysis technique for the determination the origins of meat species in a very short analysis time (30 s) without requiring complicated chromatographic, immunologic or genetic methods and instruments and also trained personnel.






