THe AtSCL26 Transcription Factor Controls Cross-talk Between GA and N Root Architecture in Arabidopsis Thaliana Roots
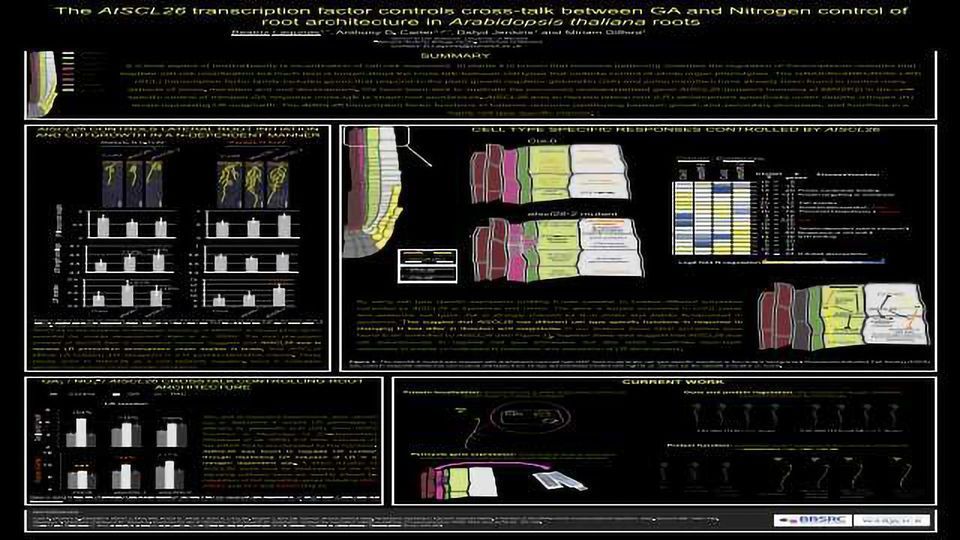
Multicellular organisms such as plants rely on highly specialised tissue-specific cell-types to perform discrete functions, and this specialisation enables modulation of plant productivity. Cell-type identity is also involved in determining which cells are capable of sensing internal and external conditions and the amplitude of downstream responses. In roots, aside from mediating growth and nutrient uptake, cell-specific transcriptional programs are fundamental to the developmental organisation of root architecture including formation of lateral branching roots that enable the root to harvest nutrients, and root nodule structures. Root nodules form to house symbiosis between legume species with soil nitrogen fixing bacteria (rhizobia). Phenotypic and molecular evidence supports the hypothesis that developmental program enabling nodule formation arose during evolution 70 MYA from a lateral root ‘blueprint’ pre-existing in all higher plants .
Understanding the conserved regulation of development has high relevance for building resilience in food security and agriculture. Molecular integrators of environmental controlling development could act across multiple pathways, thus we reasoned that analyzing Arabidopsis genes orthologous to regulators of nodulation could shed insight on control of lateral root development. This led us to the discovery that an Arabidopsis GRAS family transcription factor controls lateral root development under specific nitrogen conditions. Since AtSCL26 is the putative ortholog of the key nodulation regulator MtNSP2 in Medicago truncatula, this gene appears to be a conserved developmental regulator. We also identified effects of AtSCL26 that are specific to epidermal and cortical cells using Fluorescence Activated Cell Sorting (FACS), paralleling the cell type specific effects found for the nitrogen-regulated MtNSP2.
Understanding the conserved regulation of development has high relevance for building resilience in food security and agriculture. Molecular integrators of environmental controlling development could act across multiple pathways, thus we reasoned that analyzing Arabidopsis genes orthologous to regulators of nodulation could shed insight on control of lateral root development. This led us to the discovery that an Arabidopsis GRAS family transcription factor controls lateral root development under specific nitrogen conditions. Since AtSCL26 is the putative ortholog of the key nodulation regulator MtNSP2 in Medicago truncatula, this gene appears to be a conserved developmental regulator. We also identified effects of AtSCL26 that are specific to epidermal and cortical cells using Fluorescence Activated Cell Sorting (FACS), paralleling the cell type specific effects found for the nitrogen-regulated MtNSP2.






